Difference between revisions of "BIN-SX-Chimera"
m |
m |
||
| Line 97: | Line 97: | ||
:'''3.''' When you are done with everything, add the following category tag '''to the end of page''': | :'''3.''' When you are done with everything, add the following category tag '''to the end of page''': | ||
| − | + | ::<code><nowiki>[[Category:EVAL-SX-Chimera]]</nowiki></code>. | |
Once the page has been saved with this tag, it is considered "submitted". | Once the page has been saved with this tag, it is considered "submitted". | ||
| Line 109: | Line 109: | ||
:# There are a number of tasks in which you are explicitly asked you to submit code or other text for credit. Put all of these submission on this one page. | :# There are a number of tasks in which you are explicitly asked you to submit code or other text for credit. Put all of these submission on this one page. | ||
:# When you are done with everything, add the following category tag '''to the end of page''': | :# When you are done with everything, add the following category tag '''to the end of page''': | ||
| − | + | ::<code><nowiki>[[Category:EVAL-SX-Chimera]]</nowiki></code>. | |
Once the page has been saved with this tag, it is considered "submitted". | Once the page has been saved with this tag, it is considered "submitted". | ||
| Line 121: | Line 121: | ||
:* Create a new page on the student Wiki as a subpage of your User Page. Develop your visual analysis there. | :* Create a new page on the student Wiki as a subpage of your User Page. Develop your visual analysis there. | ||
:* When you are done with everything, add the following category tag '''to the end of page''': | :* When you are done with everything, add the following category tag '''to the end of page''': | ||
| − | + | ::<code><nowiki>[[Category:EVAL-SX-Chimera]]</nowiki></code>. | |
Once the page has been saved with this tag, it is considered "submitted". | Once the page has been saved with this tag, it is considered "submitted". | ||
Revision as of 03:21, 3 November 2018
UCSF Chimera: Structure Visualization and Analysis
(UCSF Chimera; Structure visualization; Structure analysis)
Abstract:
This unit introduces the molecular viewer UCSF Chimera, starts you off on a routine to practice stereo viewing of molecular models, and teaches how to use Chimera to solve a number of common visualization and analysis tasks.
|
Objectives:
|
Outcomes:
|
Deliverables:
- Time management: Before you begin, estimate how long it will take you to complete this unit. Then, record in your course journal: the number of hours you estimated, the number of hours you worked on the unit, and the amount of time that passed between start and completion of this unit.
- Journal: Document your progress in your Course Journal. Some tasks may ask you to include specific items in your journal. Don't overlook these.
- Insights: If you find something particularly noteworthy about this unit, make a note in your insights! page.
Prerequisites:
This unit builds on material covered in the following prerequisite units:
Contents
Evaluation
This learning unit can be evaluated for a maximum of 6 marks. If you want to submit tasks for this unit for credit you have the following options. If you have any questions about these options, discuss on the mailing list.
- Short Report option
- 1. Create a new page on the student Wiki as a subpage of your User Page.
- 2. Alpha helices orient peptide carbonyls parallel to their axis and since the C=O bond is polarized with an excess of electrons near the carboyl oxygen, this creates an induced dipole along the length of the helix, with a positive partial charge towards the beginning of the helix, and a negative partial charge towards its end. To compensate, we often find negativeley charged residues at the beginning of the helix and positively charged residues at the end - to stabilize the helix. However, for DNA binding, electrostatic complementarity would make us assume that alpha helices in DNA binding proteins are generally oriented with the (+)-dipole close to the DNA phosphates, and that there are no compensating charged residues at the termini. Write a short analysis on one of the three following topics - A, B, or C. The goal of this short report is to connect visualization to biology.
- A - Helix dipoles: Nucleosome core
- A.1 Open Chimera and load the structure 5XF3.
- A.2 Study the complex with respect to this assumption. Produce 2 or more informative stereo-images that illustrate your findings.a
- A.3 Write up, illustrate, and interpret your findings in a short report.
- B - Helix dipole: Transcription factor
- B.1 Open Chimera and load the structure 4Y60.
- B.2 Study the complex with respect to this assumption. Produce 2 or more informative stereo-images that illustrate your findings.
- B.3 Write up, illustrate and interpret your findings in a short report.
- C - Helix dipole: Transcription factor
- C.1 Open Chimera and load the structure 4ZSF.
- C.2 Study the complex with respect to this assumption. Produce 2 or more informative stereo-images that illustrate your findings.
- C.3 Write up, illustrate and interpret your findings in a short report.
- 3. When you are done with everything, add the following category tag to the end of page:
[[Category:EVAL-SX-Chimera]].
Once the page has been saved with this tag, it is considered "submitted". Do not change your submission after this tag has been added. The page will be marked and the category tag will be removed by the instructor.
- Option to produce a "protein structure gallery"" entry
- Illustrating a research article well is a crucial skill that can significantly enhance the chances of it being accpted for publication. Chimera provides great support for all sorts of visual and quantitative analysis, in this deliverable you showcase some of its options to highlight a biological fact, or illustrate the use of the Chimera program. Navigate to the Instructions and Topics page on the Student Wiki to view an example and choose a topic.
- Create a new page on the student Wiki as a subpage of your User Page. Develop your visual analysis there.
- When you are done with everything, add the following category tag to the end of page:
[[Category:EVAL-SX-Chimera]].
Once the page has been saved with this tag, it is considered "submitted". Do not change your submission after this tag has been added. The page will be marked and the category tag will be removed by the instructor.
Contents
Molecular graphics: UCSF Chimera
To view molecular structures, we need a tool to visualize the three dimensional relationships of atoms. A molecular viewer is a program that takes 3D structure data and allows you to display and explore it. For a number of reasons, I use the UCSF Chimera viewer for this course:
- Chimera is free and open;
- It creates very appealing graphics;
- It is under ongoing development and is well maintained (actually, Chimera's next generation version "ChimeraX" will be released soon);
- It provides an array of useful utilities for structure analysis; and,
- besides an intuitive, menu driven interface, Chimera can be scripted via its command line, or even programmed via its in-built python interpreter.
Chimera: First steps
Task:
- Installation
- Access the Chimera homepage and navigate to the Download section.
- Find the the newest version for your platform in the table and click on the file to download it.
- Follow the instructions to install Chimera.
- First tutorial
The Chimera User Guide site has a set of associated tutorials and there are more tutorials linked from here.
- Work through Part 1 and Part 2 of the "Getting Started" Tutorial
- There are many preference options to set, for this course open UCSF Chimera → Preferences; choose Category: Messages and set Show status line: true. This will place a status-line with three shortcut icons on the bottom of the viewer window. The status line returns item information, and the shortcut icons access the Quick Access screen (lightning bolt), the Task panel (i), and the Selection Inspector panel (magnifying glass) through which you cn set detailed display options. Then close the Preferences panel.
Stereo Viewing
Stereo viewing is easy to learn with a molecular viewer like Chimera.
Being able to visualize and experience structure in 3D is an essential skill if you are at all serious about understanding the molecules of molecular biology. This is not sufficiently realized in the field: many molecular biologists have never invested the effort it takes to learn the skill and thus will tell you that it is not actually necessary, and you can get by regardless. Of course you are talking to a biased population – unless you have experienced and worked with stereo images, you can't understand how much you are actually missing. But once you have acquired the skill, you'll regret not having been taught earlier. Speak to people who use stereo vision: seeing molecules in 3-D is like the difference between seeing a photograph of a place and actually being there. In 3D you can appreciate size, scale, distance, spatial relations all at a single glance. I insist: you can't understand structure unless you experience it in 3D.
Even though hardware devices exist that help in the three-dimensional perception of computer graphics images, for the serious structural biologist there is really no alternative to being able to fuse stereo pair images by looking at them. Chimera is an excellent tool to practice stereo viewing and develop the skill. Stereo images consist of a left-eye and a right-eye view of the same object, with a slight rotation around the vertical axis (about 5 degrees). Your brain can accurately calculate depth from these two images, if they are presented to the right and left eye separately. This means you need to look at the two images and then fuse them into a single image - this happens when the left eye looks directly at the left image and the right eye at the right image.
In this tutorial, I teach you a method to learn stereo viewing. The method is pretty foolproof - I have taught this many years in my classes with virtually 100% success rates. But I can only teach you the method – learning must be done by you.
Some people find convergent (cross-eyed) stereo viewing easier to learn. I recommend the divergent (wall-eyed) viewing - not only because it is much more comfortable in my experience, but also because it is the default way in which stereo images in books and manuscripts are presented. The method explained below will only work for learning to view divergent stereo pairs.
Physiology
In order to visually fuse stereo image pairs, you need to override a vision reflex that couples divergence and focusing. This needs to be practiced for a while. Usually 5 to 10 minutes of practice twice daily for a week should be quite sufficient. It is not as hard as learning to ride a bicycle, but you need to practice regularly for some time, maybe 10 or 20 sessions of 3 to 5 minute over a period of a week or two. Once you have acquired the skill, it is really very comfortable and can be done effortlessly and for extended periods. You will enter a new world of molecular wonders!
Practice
Task:
Here are step by step instructions of how to practice stereo viewing with Chimera.
- Load the Mbp1 APSES domain structure 1BM8 small protein into Chimera and display it as a simple backbone model.
- Start Chimera.
- Select File→Fetch by ID... choose PDB, type 1BM8 in the form and click Fetch.
- Choose Presets → Publication 1 ..., and
- ... Tools → Viewing Controls → Effects: check depth cueing ON and shadows OFF; use the Side View menu to move the clipping planes close to your model.
- Apply a nice coloring ramp: Tools → Depiction → Rainbow and click Apply
- Set the stereo display to view the scene as a side-by-side stereo pair:
- Tools → Viewing Controls → Camera and select camera mode: wall-eye stereo.
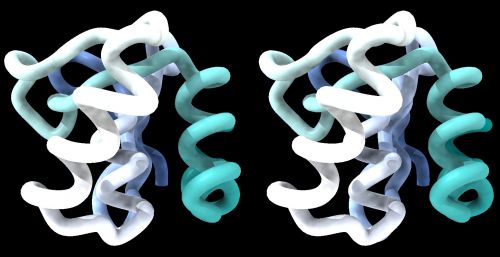
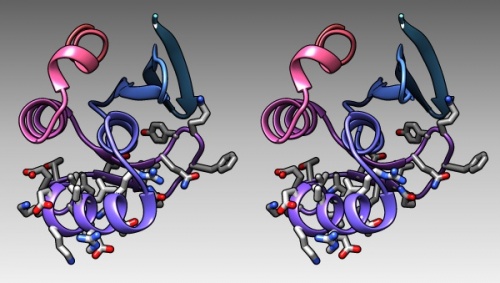
The model could look something like this:
1BM8: Mbp1 transcription factor APSES domain rendered as a ribbon model, with depth-cueing applied and a colour-ramp emphasizing the fold from N-C terminus. The molecule is shown in wall-eye stereo: the left-hand image is rotated correctly for the left eye. You should resize the window of your molecular viewer (Chimera) until equivalent points are about 15% less than your pupil separation apart.
- For the next step, you need to know how far your eyes are apart. Not just approximately, but pretty accurately. Measure the distance between your pupils in front of a mirror, or have soemone help you.
- Resize the horizontal distance of the viewing window by dragging its lower right-hand corner. This changes the separation of the two views.
- Resize the window so that two equivalent points on the protein are about 15% closer together on the screen than the pupils of your eyes are apart. Calculate, and measure eg. the separation between a protruding loop on the left and right image. Don't just guess, measure the distance, and adjust your on-screen scene to better than two or three millimetres of the correct separation.
- Also, the images themselves should be small: about the size of a postage stamp.
- Now follow these instructions exactly
- Touch your nose and forehead to the screen to get your eyes directly in front of the two images. Make sure you see the right image with your right eye, the left with your left eye. Of course, since you are so close, the images will be blurred. Nevertheless, you should see one solid, three dimensional shape in the centre, plus two peripheral images of the same view on the sides. You see three copies of the same scene, but only the fused, overlapping centre scene appears three-dimensional; the other two become less noticeable as you practice more, your brain simply begins editing them out.
- Without moving your head, resize the window slightly left and right until the centre image overlaps and "fuses". This way you find the exact distance for the images that works best for you. Slowly rotating the protein with the mouse helps generate the impression of a 3D object floating before you. Gaze at the image in the centre.
- Spend some time with this. Don't worry that it is out of focus. Imagine that you are looking at something underwater with your eyes open. But make sure that you see one central fused image and it appears three-dimensional to you. Don't continue unless you can achieve this. Ask for help if it doesn't work for you.
- Once you see the protein in 3D, try to move your head backwards slowly, until the structure comes into focus. Do not voluntarily try to focus, since this will induce your eyes to converge and you will lose the 3D effect. You should be relaxed, and passively achieve this effect. Don't force it. After a short distance, you will probably lose the 3D effect. Once you loose the 3D effect, pause, close your eyes, then look somewhere else. Relax, take a deep breath and start over with your forehaed on the screen.
- You will find that you will be able to hold the image for longer, then move your head further back, as you practice this. Give it time, you will be able to achieve focus on the #D object.
- Now start a practice routine!
- Practice this procedure patiently, three or four times daily for perhaps 3 to 5 minutes. Between lectures is ideal. Keep your scene in Chimera open and go to it frequently. But stop, when your head feels funny. Don't force yourself. Better to practice more frequently for shorter periods.
- Keep it up until you have mastered the skill.
After time and with practice, it will become easier and easier to achieve the effect. Also you will become quite independent of the distance of equivalent points, thus you can increase the viewer window size and take advantage of the increased resolution.
It might take you about a week or ten days to master this, with regular training it will become very easy. And, the best thing is, you do not easily forget this skill. It is like riding a bicycle, equalizing pressure in your ears while scuba diving, or circular breathing to play the didgeridoo: once you teach your body what to do, it remembers.
What could possibly go wrong? ... Click to expand.→
Some people get confused as to what they are "supposed" to see. You see two images of an object and you see them with each eye, i.e. in principle there are four images. The two central images (image L as seen with the left eye, image R as seen with the right eye) should overlap in the middle; these two images fuse in your visual system to create one 3D image. The two peripheral images still remain; since you don't concentrate on them, your visual system will edit them out of consciousness as you gain experience.
Chimera uses a default distance to screen that is too close and that exaggerates the depth of the scene to a degree that it is difficult to fuse the stereo pairs. Choose Tools → Viewing controls → Camera and set the distance to screen to 50 cm. This will make stereo viewing easier and will also give a better match between distance estimates in all three dimensions. Check this every time you start a new session, and whenever you find it difficult to make sense of a scene.
There are two situations which interfere with stereo viewing. One such situation is if the images presented to your eye are of unequal size. This can happen if you are using glasses with significantly different correction for each eye - the lenses then have different magnifications. The other situation is when equivalent points of the images are vertically misaligned, i.e. one of the images is shifted up or down, or rotated. This can occur when your head is tilted relative to the image. Keep your head straight[1].
Images that are difficult to see in 3D are images that are rendered differently for the left- and right view: non-aligned jagged edges, differing shadows or highlights disturb the stereo effect. To prevent this, try to apply visual effects judiciously. Chimera is quite good with it's rendering but e.g. specular highlights in some molecular viewers are quite inconsistent between left and right. Even in Chimera, hard shadows are not quite right, better to turn shadow effects off. (If you are a programmer, remember to write your code to move the camera location, don't rotate the object because that will alter which parts of the model are shadowed!)
Of course, if you are practicing wall-eyed viewing ("side-by-side"), the so-called cross-eyed stereo images won't work for you. They look like the real thing, but the left-eye view is on the right-hand side and vice versa. You can tell that they are wrong when you achieve the image fusion because the 3-D effect seems to be all wrong. This is because the image becomes inverted in depth: near points appear far away and vice versa. If you are looking at a simple line drawing, you can't tell that this is happening. However if there are any secondary depth cues in the image, such as occlusions, shadows, highlights etc., they are in the wrong places, won't work to enhance the depth effect and just generally look weird. Once you are comfortable with stereo viewing, you can try this deliberately by selecting cross-eyed display in Chimera. Inverted space is very strange.
Some more examples of stereo scenes can be found on the Stereo vision practice page on this Wiki.
Global properties
In this series of tasks we will showcase some of the globally applied tools that help us study molecular structure.
A Ramachandran plot
Task:
- To reset all views and selections, choose Tools → General Controls → Model Panel. Select the 1BM8 model (if it's still loaded from your stereo exercises) and click the close button to remove it.
- In the message bar at the bottom of the viewer window, click on the "lightning bolt" icon at the bottom (this is the "Quick Access Screen"). You should see a button labelled 1BM8 on the right. This is where you will find recent structures. Click
1BM8to re-load it[2]. - Choose Presets → Interactive 2 (all atoms) for a detailed view.
- Choose Tools → General Controls → Model Panel.
- Look for the Option Ramachandran plot... in the choices on the right.
- Click the button and study the result. The dots in this Ramachandran Plot represent the phi-psi angle combinations for residue backbones. We see that they are well distributed, this is a high-resolution structure essentially without outliers. Clicking on a dot selects a residue in the structure viewer (selected residues have a green contour).
- Choose File → Fetch by ID and fetch
1L3G, an NMR structure of the Mbp1 APSES domain. Chimera loads the 19 models that comprise this structure dataset. - In the Tools → General Controls → Model Panel, select 1BM8 and click on hide.
- Then select 1LG3 and click group/ungroup to be able to address the models individually. Select any of the models individually and click again on Ramachandran plot. You will see that the points are much more dispersed, and there are a number of outliers that have comparatively high-energy conformations.
B-factors
Task:
- Choose Tools → General Controls → Model Panel, click/drag over the 1LG3 models and click close to remove them again.
- To explore B-Factors in the 1BM8 model, click show to view it again.
- Choose Tools → Structure Analysis → Render byAttribute.
- Select Attributes of atoms, Model 1BM8 and Attribute: bfactor. A histogram appears with sliders that allow you to render the distribution of values found in the structure for this attribute.
- Let's colour the atoms by B-Factor. Click on the colours tab. A standard colouring scheme is blue - white - red, but you can move the sliders, add new thresholds, and colour them individually by clicking on the colour patch to create your own colour spectrum, e.g. from black via red to white, in a black-body spectrum. Click Apply.
- Choose Actions → Atoms/Bonds → stick to give the bonds more volume. You will find that the core of the protein has low temperature factors, and the surface has a number of "glowingly" mobile sidechains and loops.
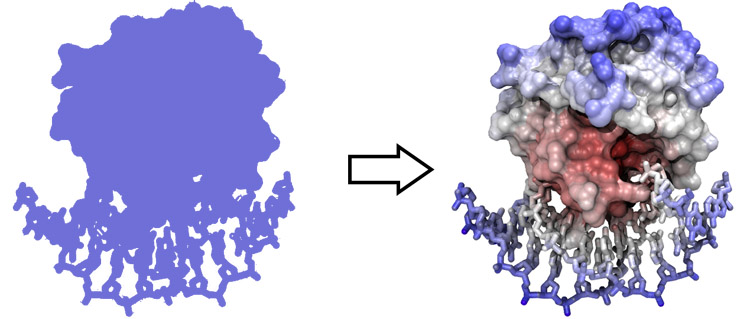
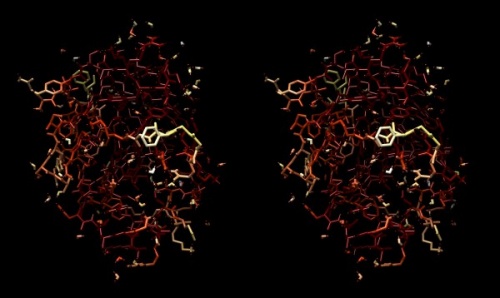
Structure of the yeast transcription factor Mbp1 DNA binding domain (1BM8) coloured by B-factor (thermal factor). The protein bonds are shown in a "stick" model, coloured with a spectrum that emulates black-body radiation. Note that the interior of the protein is less mobile, some of the surface loops are highly mobile (or statically disordered, X-ray structures can't distinguish that) and the discretely bound water molecules that are visible in this high-resolution structure are generally more mobile than the residues they bind to.
Electrostatics
Task:
- Start by getting Chimera and your model in a fresh state. Choose File → Close Session and click on the 1bm8 button to relaod the structure.
- To visualize the electrostatic potential of the protein, mapped on the surface, first select Presets → Interactive 2... and Actions → Color → cyan for a vividly contrasting color, then Select → Structure → Solvent and Actions → Atoms/Bonds → hide.
- Before mapping properties to a surface, a surface object has to be computed. Once it has been computed it will appear in the Model Panel, and in other windows that operate on surfaces, and can be separately selected and manipulated. To compute the solvent-excluded surface of the protein, simply choose Select → Structure → Protein and Actions → Surface → show.
- A simple electrostatic potential calculation just assumes Coulomb charges. A more accurate calculation of full Poisson-Boltzmann potentials is also available. Select Tools → Surface/Binding Analysis → Coulombic Surface Coloring.
- Make sure the surface object is selected in the form (it should be selected by default since there is only one surface), keep the default parameters and click Apply.
- Use Actions → Surface → Transparency → 30% to make the protein backbone somewhat visible.
- Open the Tools → Viewing Controls → Lighting window → and set mode from two-point to ambient. This reduces shadowing and reflections on the surface and thus emphasizes the color values - here our focus is not on shape, but on property.
- Use the Effects tab of the Viewing Panel to uncheck shadows (turns them off) and depth-cueing and silhouettes on. This adds visual depth cues which compensate for the loss of shape information by using a flat lighting model.
- Can you identify the putative DNA binding site from the electrostatic potential of the 1BM8 surface?
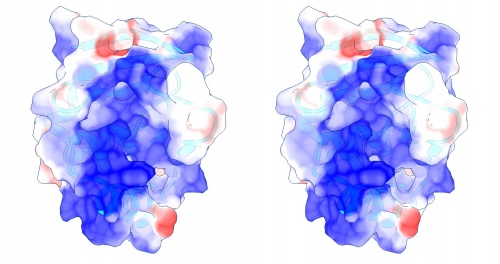
Coulomb (electrostatic) potential mapped to the solvent accessible surface of the yeast transcription factor Mbp1 DNA binding domain (1BM8). The protein backbone is visible through the transparent surface as a cartoon model, note the helix at the bottom of the structure. This helix has been suggested to play a role in forming the domain's DNA binding site and the positive (blue) electrostatic potential of the region is consistent with binding the negatively charged phosphate backbone of DNA. The other side of the domain has a negative (red) charge excess, which balances the molecule's electric charge overall, but also guides the protein-ligand interaction and supports faster on-rates.
Hydrogen bonds
Task:
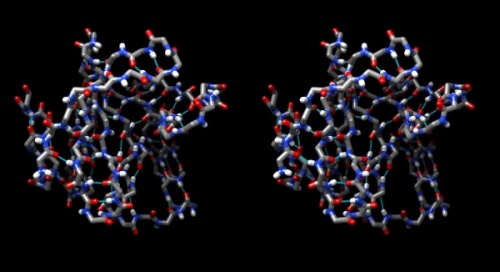
- Hydrogen bonds encode the basic folding patterns of the protein. To visualize H-bonds select Presets → Publication 1... and Actions → Color → by element.
- Use Tools → Structure Analysis → FindHBond and Apply default parameters.
- To emphasize the role of H-bonds in determining the architecture of the protein, select Select → Structure → backbone → full and then Select → Invert (all models). Now Actions → Atoms/bonds → hide will show only the backbone with its H-bonds.
Chimera sequence interface
In this task we will explore the sequence interface of Chimera, use it to select specific parts of a molecule, and colour specific regions (or residues) of a molecule separately.
Task:
- Display the protein in Presets → Interactive 1 mode and familiarize yourself with its topology of helices and strands.
- Now turn hydrogen bonds off: the menu commands of Chimera all have a command line equivalent. Open the command line by choosing Tools → General Controls → Command Line. In the Command line, type
~hbonds. The~(tilde) undoes previous commands. - Use Tools → Depiction → Rainbow to color the chain from blue to red. (You need to change the colour patches by clicking on them to open the colour editor. Choose an HSL colour model, use Saturation and Lightness 0.5 to keep the colour to somewhat subdued hues, then use the slider to choose appropriate hue values.) Click Apply.
- Open the sequence tool: Tools → Sequence → Sequence. By default, coloured rectangles overlay the secondary structure elements of the sequence.
- Hover the mouse over some residues and note that the sequence number and chain is shown at the bottom of the window.
- Click/drag one residue to select it. (Simply a click wont work, you need to drag a little bit for the selection to catch on.) Note that the residue gets a green overlay in the sequence window, and it also gets selected with a green border in the graphics window.
- In the bottom of the sequence window, there are instructions how to select (multiple) regions. Clear the selection by <control> clicking into an empty spot of the viewer. Now select the region that encompasses the residues that have been reported to form the DNA binding subdomain:
KRTRILEKEVLKETHEKVQGGFGKYQ(Taylor 2000). Show the side chains of these residues by clicking on the little green inspector icon on the viewer window, inspecting Atom and choosing displayed: true, and inspecting Bond and setting the stick radius to 0.4. - Undisplay the Hydrogen atoms by selecting the element H in the Chemistry option of the Selection Menu, and use the Action menu to hide them. Then use the effects pane of the Viewing controls menu to add silhouettes.
- Finally, give the scene a gradient grey background grey via the Actions → Color → all options... menu.
Chimera "sequence": implicit or explicit ?
You are aware of the distinction between explicit sequence (recorded in the SEQRES records of a PDB file) and implicit sequence (implied by the actual coordinates of the structure). But which of the two does the Chimera sequence window display? Let's find out.
Task:
- Open Chimera and load the 1BM8 structure from the PDB.
- Save the ccordinate file with File → Save PDB ..., use a filename of
test.pdb. - Open this file in a plain text editor: notepad, TextEdit, nano or the like - youcan use RStudio too, but not MS Word! Make sure you view the file in a fixed-width font, not proportionally spaced, i.e. Courier, not Arial. Otherwise the columns in the file won't line up.
- Find the records that begin with
SEQRES ...and confirm that the first amino acid isGLN. - Now scroll down to the
ATOMsection of the file. Identify the records for the first residue in the structure. Delete all lines for side-chain atoms except for theCBatom. This changes the coordinates for glutamine to those of alanine. - Replace the
GLNresidue name withALA(uppercase). This relabels the residue as Alanine in the coordinate section. Therefore you have changed the implicit sequence. Implicit and explicit sequence are now different. The second atom record should now look like this:
ATOM 2 CA ALA A 4 -0.575 5.127 16.398 1.00 51.22 C
- Save the file and load it in Chimera.
- Open the sequence window: does it display
QorAas the first reside?
Therefore, does Chimera use the implicit or explicit sequence in the sequence window?
Self-evaluation
Notes
- ↑ That's the same effect as when you are watching a 3D movie in the theatre, with polarized glasses: If you have weak posture or a cute neighbour and slouch to the side, your eyes become misaligned relative to the separated images on the screen, your visual system tries to compensate, but over time you get a headache. This is how stereo-haters are bred.
- ↑ If you ever get to the Quick Access Screen by mistake, just click the lightning bolt icon again to return to the Viewer.
Further reading, links and resources
- UCSF Chimera home page
- Chimera Tools Index
- Molecular Graphics Software Links– a collection of links at the PDB.
If in doubt, ask! If anything about this learning unit is not clear to you, do not proceed blindly but ask for clarification. Post your question on the course mailing list: others are likely to have similar problems. Or send an email to your instructor.
About ...
Author:
- Boris Steipe <boris.steipe@utoronto.ca>
Created:
- 2017-08-05
Modified:
- 2017-11-17
Version:
- 1.2
Version history:
- 1.2 Remove assumptions about preferences
- 1.1 Charge mapping needs to calculate the surface first
- 1.0 First live version
- 0.1 First stub - 2016 material combined
![]() This copyrighted material is licensed under a Creative Commons Attribution 4.0 International License. Follow the link to learn more.
This copyrighted material is licensed under a Creative Commons Attribution 4.0 International License. Follow the link to learn more.