Difference between revisions of "BIN-SX-Chimera"
m |
m |
||
| Line 27: | Line 27: | ||
<div id="ABC-unit-framework"> | <div id="ABC-unit-framework"> | ||
== Abstract == | == Abstract == | ||
| + | <section begin=abstract /> | ||
<!-- included from "../components/BIN-SX-Chimera.components.wtxt", section: "abstract" --> | <!-- included from "../components/BIN-SX-Chimera.components.wtxt", section: "abstract" --> | ||
... | ... | ||
| + | <section end=abstract /> | ||
{{Vspace}} | {{Vspace}} | ||
| Line 62: | Line 64: | ||
*<b>Time management</b>: Before you begin, estimate how long it will take you to complete this unit. Then, record in your course journal: the number of hours you estimated, the number of hours you worked on the unit, and the amount of time that passed between start and completion of this unit. | *<b>Time management</b>: Before you begin, estimate how long it will take you to complete this unit. Then, record in your course journal: the number of hours you estimated, the number of hours you worked on the unit, and the amount of time that passed between start and completion of this unit. | ||
<!-- included from "ABC-unit_components.wtxt", section: "deliverables-journal" --> | <!-- included from "ABC-unit_components.wtxt", section: "deliverables-journal" --> | ||
| − | *<b>Journal</b>: Document your progress in your [[FND-Journal| | + | *<b>Journal</b>: Document your progress in your [[FND-Journal|Course Journal]]. Some tasks may ask you to include specific items in your journal. Don't overlook these. |
<!-- included from "ABC-unit_components.wtxt", section: "deliverables-insights" --> | <!-- included from "ABC-unit_components.wtxt", section: "deliverables-insights" --> | ||
| − | *<b>Insights</b>: If you find something particularly noteworthy about this unit, make a note in your [[ABC-Insights|insights! page]]. | + | *<b>Insights</b>: If you find something particularly noteworthy about this unit, make a note in your [[ABC-Insights|'''insights!''' page]]. |
{{Vspace}} | {{Vspace}} | ||
Revision as of 17:32, 7 September 2017
UCSF Chimera: Structure Visualization and Analysis
Keywords: UCSF Chimera; Structure visualization; Structure analysis
Contents
This unit is under development. There is some contents here but it is incomplete and/or may change significantly: links may lead to nowhere, the contents is likely going to be rearranged, and objectives, deliverables etc. may be incomplete or missing. Do not work with this material until it is updated to "live" status.
Abstract
...
This unit ...
Prerequisites
You need to complete the following units before beginning this one:
Objectives
...
Outcomes
...
Deliverables
- Time management: Before you begin, estimate how long it will take you to complete this unit. Then, record in your course journal: the number of hours you estimated, the number of hours you worked on the unit, and the amount of time that passed between start and completion of this unit.
- Journal: Document your progress in your Course Journal. Some tasks may ask you to include specific items in your journal. Don't overlook these.
- Insights: If you find something particularly noteworthy about this unit, make a note in your insights! page.
Evaluation
Evaluation: Integrated Unit
- This unit should be submitted for evaluation for a maximum of 10 marks. Details TBD.
Contents
Molecular graphics: UCSF Chimera
To view molecular structures, we need a tool to visualize the three dimensional relationships of atoms. A molecular viewer is a program that takes 3D structure data and allows you to display and explore it. For a number of reasons, I use the UCSF Chimera viewer for this course:
- Chimera is free and open;
- It creates very appealing graphics;
- It is under ongoing development and is well maintained;
- It provides an array of useful utilities for structure analysis; and,
- besides an intuitive, menu driven interface, Chimera can be scripted via its command line, or even programmed via its in-built python interpreter.
Task:
- Access the Chimera homepage and navigate to the Download section.
- Find the the newest version for your platform in the table and click on the file to download it.
- Follow the instructions to install Chimera.
Global properties
In this series of tasks we will showcase some of the globally applied tools that help us study molecular structure.
A Ramachandran plot
Task:
- To reset all views and selections, choose Favorites → Model Panel. Select the 1BM8 model and click the close button to remove it.
- In the graphics window, click on the "lightning bolt" icon at the bottom. You should see a button labelled 1BM8 on the right. This is where you will find recent structures. Click
1BM8to re-load it. - Choose Presets → Interactive 2 (all atoms) for a detailed view.
- Choose Favorites → Model Panel
- Look for the Option Ramachandran plot... in the choices on the right.
- Click the button and study the result. The dots in thisRamachandran Plot represent the phi-psi angle combinations for residue backbones. We see that they are well distributed, this is a high-resolution structure essentially without outliers. Clicking on a dot selects a residue in the structure viewer (selected residues have a green contour).
- Choose File → Fetch by ID and fetch
1L3G, an NMR structure of the Mbp1 APSES domain. Chimera loads the 19 models that comprise this structure dataset. - In the Favorites → Model Panel, select 1BM8 and click on hide.
- Then select 1LG3 and click group/ungroup to be able to address the models individually. Select any of the models individually and click again on Ramachandran plot. You will see that the points are much more dispersed, and there are a number of outliers that have comparatively high-energy conformations.
B-factors
Task:
- Choose Favorites → Model Panel, click/drag over the 1LG3 models and click close to remove them again.
- To explore B-Factors in the 1BM8 model, click show to view it again.
- Choose Tools → Structure Analysis → Render byAttribute.
- Select Attributes of atoms, Model 1BM8 and Attribute: bfactor. A histogram appears with sliders that allow you to render the distribution of values found in the structure for this attribute.
- Let's colour the atoms by B-Factor. Click on the colours tab. A standard colouring scheme is blue - white - red, but you can move the sliders, add new thresholds, and colour them individually by clicking on the colour patch to create your own colour spectrum, e.g. from black via red to white, in a black-body spectrum. Click Apply.
- Choose Actions → Atoms/Bonds → stick to give the bonds more volume. You will find that the core of the protein has low temperature factors, and the surface has a number of highly mobile sidechains and loops.
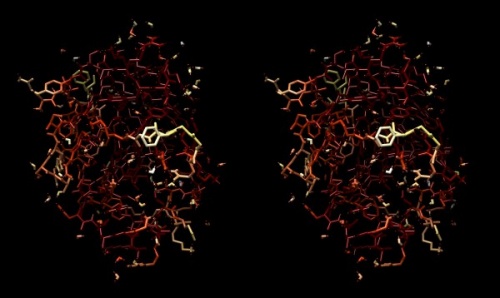
Structure of the yeast transcription factor Mbp1 DNA binding domain (1BM8) coloured by B-factor (thermal factor). The protein bonds are shown in a "stick" model, coloured with a spectrum that emulates black-body radiation. Note that the interior of the protein is less mobile, some of the surface loops are highly mobile (or statically disordered, X-ray structures can't distinguish that) and the discretely bound water molecules that are visible in this high-resolution structure are generally more mobile than the residues they bind to.
Electrostatics
Task:
- To visualize the electrostatic potential of the protein, mapped on the surface, first select Presets → Interactive 2... and Actions → Color → cyan for a vividly contrasting color.
- A simple electrostatic potential calculation just assumes Coulomb charges. A more accurate calculation of full Poisson-Boltzmann potentials is also available. Select Tools → Electrostatic/Binding Analysis → Coulombic Surface Coloring.
- Make sure the surface object is selected in the form (it should be selected by default since there is only one surface), keep the default parameters and click Apply.
- Use Actions → Surface → Transparency → 30% to make the protein backbone somewhat visible.
- Open the Tools → Viewing Controls → Lighting window → and set Intensity from two-point to ambient. This reduces shadowing and reflections on the surface and thus emphasizes the color values - here our focus is not on shape, but on property.
- Use the Effects tab to turn shadows off and depth-cueing and silhouettes on. This recreates visual cues of depth which compensate for the loss of shape information by using a flat lighting model.
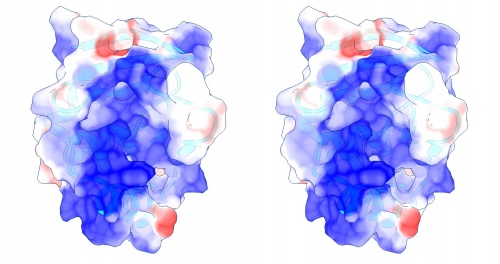
Coulomb (electrostatic) potential mapped to the solvent accessible surface of the yeast transcription factor Mbp1 DNA binding domain (1BM8). The protein backbone is visible through the transparent surface as a cartoon model, note the helix at the bottom of the structure. This helix has been suggested to play a role in forming the domain's DNA binding site and the positive (blue) electrostatic potential of the region is consistent with binding the negatively charged phosphate backbone of DNA. The other side of the domain has a negative (red) charge excess, which balances the molecule's electric charge overall, but also guides the protein-ligand interaction and supports faster on-rates.
Hydrogen bonds
Task:
- Hydrogen bonds encode the basic folding patterns of the protein. To visualize H-bonds select Presets → Publication 1... and Actions → Color → by element.
- Use Tools → Structure Analysis → FindHBond and Apply default parameters.
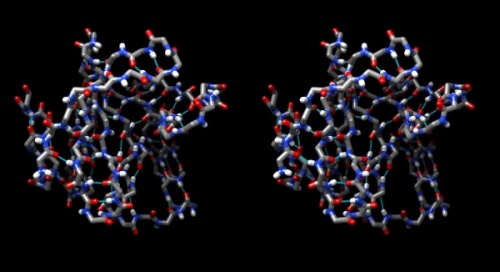
- To emphasize the role of H-bonds in determining the architecture of the protein, select Select → Structure → backbone → full and then Select → Invert (all models). Now Actions → Atoms/bonds → hide will show only the backbone with its H-bonds.
Chimera sequence interface
In this task we will explore the sequence interface of Chimera, use it to select specific parts of a molecule, and colour specific regions (or residues) of a molecule separately.
Task:
- Display the protein in Presets → Interactive 1 mode and familiarize yourself with its topology of helices and strands.
- Now turn hydrogen bonds off: the menu commands of Chimera all have a command line equivalent. Open the command line by clicking on the "computer" icon in the upper left corner of the viewer window. Then type "~hbonds". The "~" undoes previous commands.
- Use Tools → Depiction → Rainbow to color the chain from blue to red. (You need to change the colour patches by clicking on them to open the colour editor. Choose an HSL colour model, use Saturation and Lightness 0.5 to keep the colour to somewhat subdued hues, then use the slider to choose appropriate hue values.) Click Apply.
- Open the sequence tool: Tools → Sequence → Sequence. By default, coloured rectangles overlay the secondary structure elements of the sequence.
- Hover the mouse over some residues and note that the sequence number and chain is shown at the bottom of the window.
- Click/drag one residue to select it. (Simply a click wont work, you need to drag a little bit for the selection to catch on.) Note that the residue gets a green overlay in the sequence window, and it also gets selected with a green border in the graphics window.
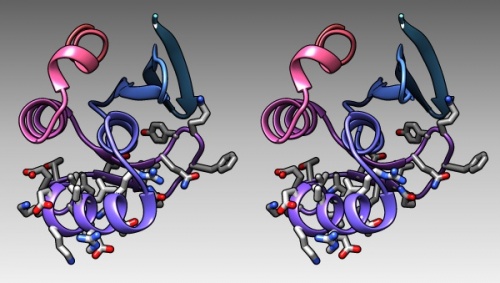
- In the bottom of the sequence window, there are instructions how to select (multiple) regions. Clear the selection by <control> clicking into an empty spot of the viewer. Now select the region that encompasses the residues that have been reported to form the DNA binding subdomain:
KRTRILEKEVLKETHEKVQGGFGKYQ(Taylor 2000). Show the side chains of these residues by clicking on the little green inspector icon on the viewer window, inspecting Atom and choosing displayed: true, and inspecting Bond and setting the stick radius to 0.4. - Undisplay the Hydrogen atoms by selecting the element H in the Chemistry option of the Selection Menu, and use the Action menu to hide them. Then use the effects pane of the Depiction menu to add a contour.
- Finally, give the scene a gradient grey background grey via the Actions → Color → all options... menu.
Chimera "sequence": implicit or explicit ?
We discussed the distinction between implicit and explicit sequence. But which one does the Chimera sequence window display? Let's find out.
Task:
- Open Chimera and load the 1BM8 structure from the PDB.
- Save the ccordinate file with File → Save PDB ..., use a filename of
test.pdb. - Open this file in a plain text editor: notepad, TextEdit, nano or the like, but not MS Word! Make sure you view the file in a fixed-width font, not proportionally spaced, i.e. Courier, not Arial. Otherwise the columns in the file won't line up.
- Find the records that begin with
SEQRES ...and confirm that the first amino acid isGLN. - Now scroll down to the
ATOMsection of the file. Identify the records for the first residue in the structure. Delete all lines for side-chain atoms except for theCBatom. This changes the coordinates for glutamine to those of alanine. - Replace the
GLNresidue name withALA(uppercase). This relabels the residue as Alanine in the coordinate section. Therefore you have changed the implicit sequence. Implicit and explicit sequence are now different. The second atom record should now look like this:
ATOM 2 CA ALA A 4 -0.575 5.127 16.398 1.00 51.22 C
- Save the file and load it in Chimera.
- Open the sequence window: does it display
QorAas the first reside?
Therefore, does Chimera use the implicit or explicit sequence in the sequence window?
Further reading, links and resources
- Chimera page
- Chimera Tools Index
- Molecular Graphics Software Links– a collection of links at the PDB.
| Taylor et al. (2000) Characterization of the DNA-binding domains from the yeast cell-cycle transcription factors Mbp1 and Swi4. Biochemistry 39:3943-54. (pmid: 10747782) |
|
[ PubMed ] [ DOI ] The minimal DNA-binding domains of the Saccharomyces cerevisiae transcription factors Mbp1 and Swi4 have been identified and their DNA binding properties have been investigated by a combination of methods. An approximately 100 residue region of sequence homology at the N-termini of Mbp1 and Swi4 is necessary but not sufficient for full DNA binding activity. Unexpectedly, nonconserved residues C-terminal to the core domain are essential for DNA binding. Proteolysis of Mbp1 and Swi4 DNA-protein complexes has revealed the extent of these sequences, and C-terminally extended molecules with substantially enhanced DNA binding activity compared to the core domains alone have been produced. The extended Mbp1 and Swi4 proteins bind to their cognate sites with similar affinity [K(A) approximately (1-4) x 10(6) M(-)(1)] and with a 1:1 stoichiometry. However, alanine substitution of two lysine residues (116 and 122) within the C-terminal extension (tail) of Mbp1 considerably reduces the apparent affinity for an MCB (MluI cell-cycle box) containing oligonucleotide. Both Mbp1 and Swi4 are specific for their cognate sites with respect to nonspecific DNA but exhibit similar affinities for the SCB (Swi4/Swi6 cell-cycle box) and MCB consensus elements. Circular dichroism and (1)H NMR spectroscopy reveal that complex formation results in substantial perturbations of base stacking interactions upon DNA binding. These are localized to a central 5'-d(C-A/G-CG)-3' region common to both MCB and SCB sequences consistent with the observed pattern of specificity. Changes in the backbone amide proton and nitrogen chemical shifts upon DNA binding have enabled us to experimentally define a DNA-binding surface on the core N-terminal domain of Mbp1 that is associated with a putative winged helix-turn-helix motif. Furthermore, significant chemical shift differences occur within the C-terminal tail of Mbp1, supporting the notion of two structurally distinct DNA-binding regions within these proteins. |
Notes
Self-evaluation
If in doubt, ask! If anything about this learning unit is not clear to you, do not proceed blindly but ask for clarification. Post your question on the course mailing list: others are likely to have similar problems. Or send an email to your instructor.
About ...
Author:
- Boris Steipe <boris.steipe@utoronto.ca>
Created:
- 2017-08-05
Modified:
- 2017-08-05
Version:
- 0.1
Version history:
- 0.1 First stub
![]() This copyrighted material is licensed under a Creative Commons Attribution 4.0 International License. Follow the link to learn more.
This copyrighted material is licensed under a Creative Commons Attribution 4.0 International License. Follow the link to learn more.