Difference between revisions of "BIN-PHYLO-Tree analysis"
m |
m |
||
| Line 41: | Line 41: | ||
<b>Deliverables:</b><br /> | <b>Deliverables:</b><br /> | ||
<section begin=deliverables /> | <section begin=deliverables /> | ||
| − | |||
<li><b>Time management</b>: Before you begin, estimate how long it will take you to complete this unit. Then, record in your course journal: the number of hours you estimated, the number of hours you worked on the unit, and the amount of time that passed between start and completion of this unit.</li> | <li><b>Time management</b>: Before you begin, estimate how long it will take you to complete this unit. Then, record in your course journal: the number of hours you estimated, the number of hours you worked on the unit, and the amount of time that passed between start and completion of this unit.</li> | ||
| − | |||
<li><b>Journal</b>: Document your progress in your [[FND-Journal|Course Journal]]. Some tasks may ask you to include specific items in your journal. Don't overlook these.</li> | <li><b>Journal</b>: Document your progress in your [[FND-Journal|Course Journal]]. Some tasks may ask you to include specific items in your journal. Don't overlook these.</li> | ||
| − | |||
<li><b>Insights</b>: If you find something particularly noteworthy about this unit, make a note in your [[ABC-Insights|'''insights!''' page]].</li> | <li><b>Insights</b>: If you find something particularly noteworthy about this unit, make a note in your [[ABC-Insights|'''insights!''' page]].</li> | ||
<section end=deliverables /> | <section end=deliverables /> | ||
| Line 52: | Line 49: | ||
<section begin=prerequisites /> | <section begin=prerequisites /> | ||
<b>Prerequisites:</b><br /> | <b>Prerequisites:</b><br /> | ||
| − | |||
This unit builds on material covered in the following prerequisite units:<br /> | This unit builds on material covered in the following prerequisite units:<br /> | ||
*[[BIN-PHYLO-Tree_building|BIN-PHYLO-Tree_building (Building Phylogenetic Trees)]] | *[[BIN-PHYLO-Tree_building|BIN-PHYLO-Tree_building (Building Phylogenetic Trees)]] | ||
| Line 72: | Line 68: | ||
| + | === Evaluation === | ||
| + | <b>Evaluation: NA</b><br /> | ||
| + | <div style="margin-left: 2rem;">This unit is not evaluated for course marks.</div> | ||
== Contents == | == Contents == | ||
| − | |||
| Line 349: | Line 347: | ||
--> | --> | ||
| − | |||
| − | |||
| − | |||
| − | |||
== Further reading, links and resources == | == Further reading, links and resources == | ||
| Line 361: | Line 355: | ||
Also: [http://www.nature.com/scitable/topicpage/reading-a-phylogenetic-tree-the-meaning-of-41956 Nature-Scitable (2008): '''Reading a Phylogenetic Tree: The Meaning of Monophyletic Groups'''] | Also: [http://www.nature.com/scitable/topicpage/reading-a-phylogenetic-tree-the-meaning-of-41956 Nature-Scitable (2008): '''Reading a Phylogenetic Tree: The Meaning of Monophyletic Groups'''] | ||
| + | == Notes == | ||
| + | <references /> | ||
{{Vspace}} | {{Vspace}} | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
<div class="about"> | <div class="about"> | ||
| Line 392: | Line 376: | ||
*0.1 First stub | *0.1 First stub | ||
</div> | </div> | ||
| − | |||
| − | |||
{{CC-BY}} | {{CC-BY}} | ||
| + | [[Category:ABC-units]] | ||
| + | {{UNIT}} | ||
| + | {{REVISE}} | ||
</div> | </div> | ||
<!-- [END] --> | <!-- [END] --> | ||
Revision as of 09:26, 25 September 2020
Analysing Phylogenetic Trees
(Species trees, gene trees and the importance of naming, Speciation and duplication signatures)
Abstract:
The analysis of mixed gene trees.
|
Objectives:
|
Outcomes:
|
Deliverables:
Prerequisites:
This unit builds on material covered in the following prerequisite units:
Contents
Evaluation
Evaluation: NA
Contents
Task:
- Read the introductory notes on analysing phylogenetic trees.
Analysis
Analysing your tree
In order to analyse your tree, you need a species tree as reference. This really is an absolute prerequisite to make your expectations about the observed tree explicit. Fortunately we have all species nicely documented in our database.
The reference species tree
Task:
To get a species tree, we make use of the smart and useful PhyloT service:
- Navigate to the PhyloT
- Execute the following R command to create a list of all taxonomy records for the species in your database (plus E. coli):
cat(paste(c(myDB$taxonomy$ID, "83333"), collapse=", "))
- Copy this list and paste it into the search field of the PhyloT page. Select: Scientific names; Internal Nodes collapsed; polytomy no; Format newick; Filename fungiTree.txt. Click on Generate tree. The file fungiTree.txt will be downloaded to your computer into your default download directory. Move it to your project directory. Then click on Visualize in iTOL and confirm the tree: the resulting tree should have twelve species names listed - ten "reference" fungi, E. coli (as the outgroup), and MYSPE. Make sure MYSPE is included! If it's not there, you did something wrong that needs to be fixed.
- Open fungiTree.txt in RStudio. This is a tree, specified in the so-called "Newick format". The topology of the tree is defined through the brackets, and the branch-lengths are all the same: this is a cladogram, not a phylogram.
Let's continue the analysis in R.
Task:
- Open RStudio and load the
ABC-unitsR project. If you have loaded it before, choose File → Recent projects → ABC-Units. If you have not loaded it before, follow the instructions in the RPR-Introduction unit. - Choose Tools → Version Control → Pull Branches to fetch the most recent version of the project from its GitHub repository with all changes and bug fixes included.
- Type
init()if requested. - Open the file
BIN-PHYLO-Tree_analysis.Rand follow the instructions.
Note: take care that you understand all of the code in the script. Evaluation in this course is cumulative and you may be asked to explain any part of code.
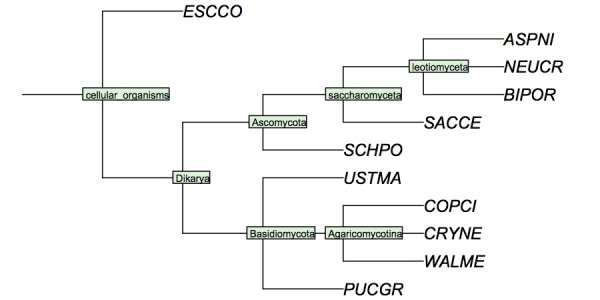
I have constructed a cladogram for many of the species we are analysing, based on data published for 1551 fungal ribosomal sequences. The six reference species are included. Such reference trees from rRNA data are a standard method of phylogenetic analysis, supported by the assumption that rRNA sequences are monophyletic and have evolved under comparable selective pressure in all species.
Cladogram of the "reference" fungi studied in the assignments. This cladogram is based on a tree returned by the NCBI Common Tree. It is thus a digest of cladistic relationships, not a representation of a specific molecular phylogeny.
Alternatively, you can look up your species in the latest version of the species tree for the fungi and add it to the tree by hand while resolving the trifurcations. See:
| Ebersberger et al. (2012) A consistent phylogenetic backbone for the fungi. Mol Biol Evol 29:1319-34. (pmid: 22114356) |
|
[ PubMed ] [ DOI ] The kingdom of fungi provides model organisms for biotechnology, cell biology, genetics, and life sciences in general. Only when their phylogenetic relationships are stably resolved, can individual results from fungal research be integrated into a holistic picture of biology. However, and despite recent progress, many deep relationships within the fungi remain unclear. Here, we present the first phylogenomic study of an entire eukaryotic kingdom that uses a consistency criterion to strengthen phylogenetic conclusions. We reason that branches (splits) recovered with independent data and different tree reconstruction methods are likely to reflect true evolutionary relationships. Two complementary phylogenomic data sets based on 99 fungal genomes and 109 fungal expressed sequence tag (EST) sets analyzed with four different tree reconstruction methods shed light from different angles on the fungal tree of life. Eleven additional data sets address specifically the phylogenetic position of Blastocladiomycota, Ustilaginomycotina, and Dothideomycetes, respectively. The combined evidence from the resulting trees supports the deep-level stability of the fungal groups toward a comprehensive natural system of the fungi. In addition, our analysis reveals methodologically interesting aspects. Enrichment for EST encoded data-a common practice in phylogenomic analyses-introduces a strong bias toward slowly evolving and functionally correlated genes. Consequently, the generalization of phylogenomic data sets as collections of randomly selected genes cannot be taken for granted. A thorough characterization of the data to assess possible influences on the tree reconstruction should therefore become a standard in phylogenomic analyses. |
Task:
- Return to the RStudio project and continue with the script to its end. Note the deliverable at the end: to print out your trees and bring them to class.
Further reading, links and resources
| Szöllősi et al. (2015) Genome-scale phylogenetic analysis finds extensive gene transfer among fungi. Philos Trans R Soc Lond., B, Biol Sci 370:20140335. (pmid: 26323765) |
|
[ PubMed ] [ DOI ] Although the role of lateral gene transfer is well recognized in the evolution of bacteria, it is generally assumed that it has had less influence among eukaryotes. To explore this hypothesis, we compare the dynamics of genome evolution in two groups of organisms: cyanobacteria and fungi. Ancestral genomes are inferred in both clades using two types of methods: first, Count, a gene tree unaware method that models gene duplications, gains and losses to explain the observed numbers of genes present in a genome; second, ALE, a more recent gene tree-aware method that reconciles gene trees with a species tree using a model of gene duplication, loss and transfer. We compare their merits and their ability to quantify the role of transfers, and assess the impact of taxonomic sampling on their inferences. We present what we believe is compelling evidence that gene transfer plays a significant role in the evolution of fungi. |
| Ebersberger et al. (2012) A consistent phylogenetic backbone for the fungi. Mol Biol Evol 29:1319-34. (pmid: 22114356) |
|
[ PubMed ] [ DOI ] The kingdom of fungi provides model organisms for biotechnology, cell biology, genetics, and life sciences in general. Only when their phylogenetic relationships are stably resolved, can individual results from fungal research be integrated into a holistic picture of biology. However, and despite recent progress, many deep relationships within the fungi remain unclear. Here, we present the first phylogenomic study of an entire eukaryotic kingdom that uses a consistency criterion to strengthen phylogenetic conclusions. We reason that branches (splits) recovered with independent data and different tree reconstruction methods are likely to reflect true evolutionary relationships. Two complementary phylogenomic data sets based on 99 fungal genomes and 109 fungal expressed sequence tag (EST) sets analyzed with four different tree reconstruction methods shed light from different angles on the fungal tree of life. Eleven additional data sets address specifically the phylogenetic position of Blastocladiomycota, Ustilaginomycotina, and Dothideomycetes, respectively. The combined evidence from the resulting trees supports the deep-level stability of the fungal groups toward a comprehensive natural system of the fungi. In addition, our analysis reveals methodologically interesting aspects. Enrichment for EST encoded data-a common practice in phylogenomic analyses-introduces a strong bias toward slowly evolving and functionally correlated genes. Consequently, the generalization of phylogenomic data sets as collections of randomly selected genes cannot be taken for granted. A thorough characterization of the data to assess possible influences on the tree reconstruction should therefore become a standard in phylogenomic analyses. |
| Marcet-Houben & Gabaldón (2009) The tree versus the forest: the fungal tree of life and the topological diversity within the yeast phylome. PLoS ONE 4:e4357. (pmid: 19190756) |
|
[ PubMed ] [ DOI ] A recurrent topic in phylogenomics is the combination of various sequence alignments to reconstruct a tree that describes the evolutionary relationships within a group of species. However, such approach has been criticized for not being able to properly represent the topological diversity found among gene trees. To evaluate the representativeness of species trees based on concatenated alignments, we reconstruct several fungal species trees and compare them with the complete collection of phylogenies of genes encoded in the Saccharomyces cerevisiae genome. We found that, despite high levels of among-gene topological variation, the species trees do represent widely supported phylogenetic relationships. Most topological discrepancies between gene and species trees are concentrated in certain conflicting nodes. We propose to map such information on the species tree so that it accounts for the levels of congruence across the genome. We identified the lack of sufficient accuracy of current alignment and phylogenetic methods as an important source for the topological diversity encountered among gene trees. Finally, we discuss the implications of the high levels of topological variation for phylogeny-based orthology prediction strategies. |
Also: Nature-Scitable (2008): Reading a Phylogenetic Tree: The Meaning of Monophyletic Groups
Notes
About ...
Author:
- Boris Steipe <boris.steipe@utoronto.ca>
Created:
- 2017-08-05
Modified:
- 2017-10-31
Version:
- 1.0
Version history:
- 1.0 First live version.
- 0.1 First stub
![]() This copyrighted material is licensed under a Creative Commons Attribution 4.0 International License. Follow the link to learn more.
This copyrighted material is licensed under a Creative Commons Attribution 4.0 International License. Follow the link to learn more.