BIN-ALI-BLAST
BLAST heuristic sequence alignment
Keywords: BLAST algorithm and Web interface, interpretation of BLAST alignments
Contents
This unit is under development. There is some contents here but it is incomplete and/or may change significantly: links may lead to nowhere, the contents is likely going to be rearranged, and objectives, deliverables etc. may be incomplete or missing. Do not work with this material until it is updated to "live" status.
Abstract
...
This unit ...
Prerequisites
You need to complete the following units before beginning this one:
Objectives
...
Outcomes
...
Deliverables
- Time management: Before you begin, estimate how long it will take you to complete this unit. Then, record in your course journal: the number of hours you estimated, the number of hours you worked on the unit, and the amount of time that passed between start and completion of this unit.
- Journal: Document your progress in your Course Journal. Some tasks may ask you to include specific items in your journal. Don't overlook these.
- Insights: If you find something particularly noteworthy about this unit, make a note in your insights! page.
Evaluation
Evaluation: NA
- This unit is not evaluated for course marks.
Contents
Heuristic pairwise alignments: BLAST
BLAST is by a margin the most important computational tool of molecular biology. It is so important, that we have already used BLAST in the BIN-Storing_data unit, to find the most similar sequence to MBP1_SACCE in MYSPE.
Task:
- Read the introductory notes on the BLAST algorithm for fast, heuristic, database-scale sequence alignment to discover homologues.
In this unit we will use BLAST to perform Reciprocal Best Matches.
One of the important questions of model-organism based inference is: which genes perform the same function in two different organisms. In the absence of other information, our best guess is that these are the two genes that are mutually most similar. The keyword here is mutually. If MBP1_SACCE from S. cerevisiae is the best match to RES2_SCHPO in S. pombe, the two proteins are only mutually most similar if RES2_SCHPO is more similar to MBP1_SACCE than to any other S. cerevisiae protein. We call this a Reciprocal Best Match, or "RBM"[1].
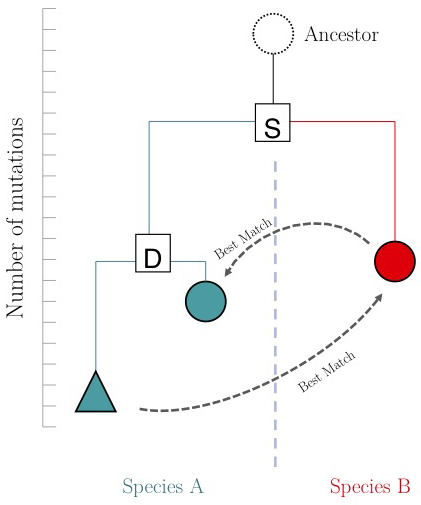
The argument is summarized in the figure on the right: genes that evolve under continuos selective pressure on their function have relatively lower mutation rates and are thus more similar to each other, than genes that undergo neo- or sub-functionalization after duplication.
However, there is a catch: proteins are often composed of multiple domains that implement distinct roles of their function. Under the assumptions above we could hypothesize:
- a gene in MYSPE that has the "same" function as the Mbp1 cell-cycle checkpoint switch in yeast should be an RBM to Mbp1;
- a gene that binds to the same DNA sites as Mbp1 should have a DNA-binding domain that is an RBM to the DNA binding domain of Mbp1.
Thus we'll compare RBMs in MYSPE for full-length Mbp1_SACCE and its DNA-binding domain, and see if the results are the same.
Full-length RBM
You have already performed the first half of the experiment: matching from S. cerevisiae to MYSPE. The backward match is simple.
Task:
- Access BLAST and follow the link to the protein blast program.
- Enter the RefSeq ID for
MBP1_MYSPEin the Query sequence field. - Select
refseq_proteinas the database to search in, and enterSaccharomyces cerevisiae (taxid:4932)to restrict the organism for which hits are reported. - Run BLAST. Examine the results.
If your top-hit is NP_010227, you have confirmed the RBM between Mbp1_SACCE and Mbp1_MYSPE. If it is not, let me know. I expect this to be the same and would like to verify your results if it is not[2].
RBM for the DNA binding domain
The DNA-binding domain of Mbp1_SACCE is called an APSES domain. If the RBM between Saccharomyces cerevisiae Mbp1 and MYSPE is truly an orthologue, we expect all of the protein's respective domains to have the RBM property as well. But let's not simply assume what we can easily test. We'll define the sequence of the APSES domain in MBP1_SACCE and MYSPE and see how these definitions reflect in a BLAST search.
Defining the range of the APSES domain annotation
The APSES domain is a well-defined type of DNA-binding domain that is ubiquitous in fungi and unique in that kingdom. Structurally it is a member of the Winged Helix-Turn-Helix family. Recently it was found that it is homologous to the somewhat shorter, prokaryotic KilA-N domain; thus the APSES domain was retired from pFam and instances were merged into the KilA-N family. However InterPro has a KilA-N entry but still recognizes the APSES domain.
KilA-N domain boundaries in Mbp1 can be derived from the results of a CDD search with the ID 1BM8_A (the Mbp1 DNA binding domain crystal structure). The KilA-N superfamily domain alignment is returned.
- (pfam 04383): KilA-N domain; The amino-terminal module of the D6R/N1R proteins defines a novel, conserved DNA-binding domain (the KilA-N domain) that is found in a wide range of proteins of large bacterial and eukaryotic DNA viruses. The KilA-N domain family also includes the previously defined APSES domain. The KilA-N and APSES domains may also share a common fold with the nucleic acid-binding modules of the LAGLIDADG nucleases and the amino-terminal domains of the tRNA endonuclease.
10 20 30 40 50 60 70 80
....*....|....*....|....*....|....*....|....*....|....*....|....*....|....*....|
1BM8A 16 IHSTGSIMKRKKDDWVNATHILKAANFAKaKRTRILEKEVLKETHEKVQ---------------GGFGKYQGTWVPLNIA 80
Cdd:pfam04383 3 YNDFEIIIRRDKDGYINATKLCKAAGETK-RFRNWLRLESTKELIEELSeennvdkseiiigrkGKNGRLQGTYVHPDLA 81
90
....*....|....
1BM8A 81 KQLA----EKFSVY 90
Cdd:pfam04383 82 LAIAswisPEFALK 95
Note that CDD and SMART are not consistent in how they apply pFam 04383 to the Mbp1 sequence. See annotation below.
The CDD KilA-N domain definition begins at position 16 of the 1BM8 sequence. But virtually all fungal APSES domains have a longer, structurally defined, conserved N-terminus. Blindly applying the KilA-N domain definition to these proteins would lose important information. For most purposes we will prefer the sequence spanned by the 1BM8_A structure. The sequence is given below, the KilA-N domain is coloured dark green. By this definition the APSES domain is 99 amino acids long and comprises residues 4 to 102 of the NP_010227 sequence.
10 20 30 40 50 60 70 80
....*....|....*....|....*....|....*....|....*....|....*....|....*....|....*....|
1BM8A 1 QIYSARYSGVDVYEFIHSTGSIMKRKKDDWVNATHILKAANFAKAKRTRILEKEVLKETHEKVQGGFGKYQGTWVPLNIA 80
90
....*....|....*....
1BM8A 81 KQLAEKFSVYDQLKPLFDF 99
- Yeast APSES domain sequence in FASTA format
>APSES_MBP1 Residues 4-102 of S. cerevisiae Mbp1 QIYSARYSGVDVYEFIHSTGSIMKRKKDDWVNATHILKAANFAKAKRTRI LEKEVLKETHEKVQGGFGKYQGTWVPLNIAKQLAEKFSVYDQLKPLFDF
- Synopsis of ranges
| Domain | Link | Length | Boundary | Range (Mbp1) | Range (1BM8) |
| KilA-N: pfam04383 (CDD) | CDD alignment | 72 | STGSI ... KFSVY | 21 - 93 | 18 - 90 |
| KilA-N: pfam04383 (SMART) | Smart main page | 79 | IHSTG ... YDQLK | 19 - 97 | 16 - 94 |
| KilA-N: SM01252 (SMART) | Smart main page | 84 | TGSIM ... DFTQT | 22 - 105 | 19 - 99... |
| APSES: Interpro IPR003163 | (Interpro) | 130 | QIYSA ... IRSAS | 3 - 133 | 1 - 99... |
| APSES (1BM8) | – | 99 | QIYSA ... PLFDF | 4 - 102 | 1 - 99 |
Executing the forward search
Task:
- Access BLAST and follow the link to the protein blast program.
- Forward search:
- Paste only the APSES domain sequence for
MBP1_SACCEin the Query sequence field (copy the sequence from above). - Select
refseq_proteinas the database to search in, and enter the correct taxonomy ID for MYSPE in the Organism field. - Run BLAST. Examine the results.
- If the top hit is the same protein you have already seen, oK. If it's not add it to your protein database in RStudio.
- Paste only the APSES domain sequence for
With this we have confirmed the sequence with the most highly conserved APSES domain in MYSPE. Can we take the sequence for the reverse search from the alignment that BLAST returns? Actually, that is not a good idea. The BLAST alignment is not guaranteed to be optimal. We should do an optimal sequnece alignment instead. That is: we use two different tools here for two different purposes: we use BLAST to identify one protein as the most similar among many alternatives and we use optimal sequence alignment to determine the best alignment between two sequences. That best alignment is what we will annotate as the MYSPE APSES domain.
Alignment to define the MYSPE APSES domain for the reverse search
Task:
- Return to your RStudio session.
- Study and work through the code in the
APSES Domain annotation by alignmentsection of theBCH441_A04.Rscript
Executing the reverse search
Task:
- Paste the the APSES domain sequence for the MYSPE best-match and enter it into Query sequence field of the BLAST form.
- Select
refseq_proteinas the database to search in, and enterSaccharomyces cerevisiae (taxid:4932)to restrict the organism for which hits are reported. - Run BLAST. Examine the results.
- Select
If your top-hit is again NP_010227, you have confirmed the RBM between the APSES domain of Mbp1_SACCE and Mbp1_MYSPE. If it is not, let me know. There may be some organisms for which the full-length and APSES RBMs are different and I would like to discuss these cases.
Further reading, links and resources
Notes
- ↑ Note that RBMs are usually orthologues, but the definition of orthologue and RBM is not the same. Most importantly, many orthologues are not RBMs. We will explore this more when we discuss phylogenetic inference.
- ↑ One such case we encountered involved a protein that has a corrupted annotation for the DNA binding domain. It appears to be the correct orthologue, but it only has the second highest BLAST score.
Self-evaluation
If in doubt, ask! If anything about this learning unit is not clear to you, do not proceed blindly but ask for clarification. Post your question on the course mailing list: others are likely to have similar problems. Or send an email to your instructor.
About ...
Author:
- Boris Steipe <boris.steipe@utoronto.ca>
Created:
- 2017-08-05
Modified:
- 2017-08-05
Version:
- 0.1
Version history:
- 0.1 First stub
![]() This copyrighted material is licensed under a Creative Commons Attribution 4.0 International License. Follow the link to learn more.
This copyrighted material is licensed under a Creative Commons Attribution 4.0 International License. Follow the link to learn more.