BIO Assignment Week 8
Assignment for Week 7
Predictions: Homology Modeling
| < Assignment 6 | Assignment 8 > |
Note! This assignment is currently active. All significant changes will be announced on the mailing list.
Concepts and activities (and reading, if applicable) for this assignment will be topics on the next quiz.
Contents
Introduction
In order to understand how specific residues in the sequence contribute to the putative function of the protein, and why and how they are conserved throughout evolution, we would need to study an explicit molecular model of an APSES domain protein, bound to its cognate DNA sequence. Explanations of a protein's observed properties and functions can't rely on the general fact that it binds DNA, we need to consider details in terms of specific residues and their spatial arrangement. In particular, it would be interesting to correlate the conservation patterns of key residues with their potential to make specific DNA binding interactions. Unfortunately, the experimental evidence we have considered in Assignment 2 (Taylor et al., 2000) is not sufficient to unambiguously define the details of how a DNA double helix might be bound. Moreover, at least two distinct modes of DNA binding are known for proteins of the winged-helix superfamily, of which the APSES domain is a member.
In this assignment you will (1) construct a molecular model of the APSES domain from the Mbp1 RBM orthologue in your assigned species.
For the following, please remember the following terminology:
- Target
- The protein that you are planning to model.
- Template
- The protein whose structure you are using as a guide to build the model.
- Model
- The structure that results from the modelling process. It has the Target sequence and is similar to the Template structure.
A brief overview article on the construction and use of homology models is linked to the resource section at the bottom of this page. That section also contains links to other sites and resources you might find useful or interesting.
A Point Mutation
To illustrate how homology modelling works in principle, let's consider changing the sequence of a single amino acid, based on a structural template.
Such minimal changes to structure models can be done directly in Chimera. Let us consider the residue A 42 of the 1BM8 structure. It is oriented towards the core of the protein, but most other Mbp1 orthologs have a larger amino acid in this position, V, or even I.
Task:
- Open
1BM8in Chimera, hide the ribbons and show all atoms as a stick model. - Color the protein white.
- Open the sequence window and select
A 42. Color it red. Choose Actions → Set pivot. Then study how nicely the alanine sidechain fits into the cavity formed by its surrounding residues. - To emphasize this better, hide the solvent molecules and select only the protein atoms. Display them as a sphere model to better appreciate the packing, i.e. the Van der Waals contacts we discussed in class. Use the Favorites → Side view panel to move the clipping plane and see a section through the protein. Study the packing, in particular, note that the additional methyl groups of a valine or isoleucine would not have enough space in the structure. Then restore the clipping planes so you can see the whole molecule.
- Lets simplify the view: choose Actions → Atoms/Bonds → backbone only → chain trace. Then select
A 42again in the sequence window and choose Actions → Atoms/Bonds → show. - Add the surrounding residues: choose Select → Zone.... In the window, see that the box is checked that selects all atoms at a distance of less then 5Å to the current selection, and check the lower box to select the whole residue of any atom that matches the distance cutoff criterion. Click OK and choose Actions → Atoms/Bonds → show.
- Select
A 42again: left-click (control click) on any atom of the alanine to select the atom, then up-arrow to select the entire residue. Now let's mutate this residue to isoleucine. - Choose Tools → Structure Editing → Rotamers and select
ILEas the rotamer type. Click OK, a window will pop up that shows you the possible rotamers for isoleucine together with their database-derived probabilities; you can select them in the window and cycle through them with your arrow keys. But note that the probabilities are very different - and thus show you high-energy and low-energy rotamers to choose from. Therefore, unless you have compelling reasons to do otherwise, try to find the highest-probability rotamer that may fit. This is where your stereo viewing practice becomes important, if not essential. It is really, really hard to do this reasonably in a 2D image! It becomes quite obvious in 3D. Btw: I find such "quantitative" work - where the real distances are important - easier in orthographic than in perspective view (cf. the Camera panel). - I find that the first rotamer is actually not such a bad fit. The
CDatom comes close to the sidechains ofI 25andL 96. But we can assume that these are somewhat mobile and can accommodate a denser packing, because - as you can easily verify in your Jalview alignment - it is NOT the case that sequences that haveI 42, have a smaller residue in position25and/or96. So let's accept the most frequentILErotamer by selecting it in the rotamer window and clicking OK (while existing side chain(s): replace is selected). - Done.
If you want to go over this in more detail, check the video tutorial on YouTube published by the NIAID bioinformatics group here. I would also encourage you to go over Part 2 of the video tutorial that discusses how to check for and resolve (by energy minimization) steric clashes. But do remember that it is not clear whether energy minimization will make your structure more correct in the sense of a smaller overall RMSD with the real, mutated protein.
What we have done here with one residue is exactly the way homology modeling works with entire sequences. The homology modelling program simply changes all amino acids to the residues of the target sequence, based on the template structure. Let's now build a homology model for YFO Mbp1.
Preparation
- We need to define our Target sequence;
- find a suitable structural Template; and
- build a Model.
Target sequence
We have encountered the PDB 1BM8 structure before, the APSES domain of saccharomyces cerevisiae Mbp1. This is a useful template to model the DNA binding domain of your RBM match. But what exactly is the aligned region of the APSES domain? We could use several approaches to define the APSES domain:
- we could use the biostrings package to calculate a pairwise sequence alignment with the
1BM8sequence, like we did previously for the full-length sequences. This would give us the domain boundaries. - we could calculate a multiple sequence alignment, while including the
1BM8sequence. This would also allow us to infer domain boundaries, actually in all sequences in our database at once. But we have found previously that such multiple sequence alignments are quite sensitive to un-alignable regions of which we have quite a few in the full length sequences. We do need an MSA, but we do need to restrict the length of the sequences we align to a reasonable region. - we could access the domain annotations at CDD or at the SMART Database, but both have interfaces that are difficult to use computationally, and have other issues: NCBI does not recognize APSES domains, only the smaller KilA-N domain, and SMART does not find APSES domains in many of our sequences.
- In our case it seems the best results are had when searching the Prosite database with the ScanProsite interface.
Task:
Let's have a first look at ScanProsite, using the yeast Mbp1 sequence. We need the UniProt ID to search Prosite. With your protein database loaded in a fresh R session, type
# (commands indented, to align their components and
# help you understand their relationship)
refDB$protein$uniProtID
which(refDB$protein$name == "MBP1")
refDB$protein$uniProtID[which(refDB$protein$name == "MBP1")]
uID <- refDB$protein$uniProtID[which(refDB$protein$name == "MBP1")]
uID- Navigate to ScanProsite, paste the UniprotID for yeast Mbp1 into the text field, select Table output for STEP 3, and START THE SCAN.
You should see four feature hits: the APSES domain, and three ankyrin domain sequences that partially overlap. We could copy and paste the start and end numbers and IDs but that would be lame. Let's get them directly from Prosite instead, because we will want to fetch a few of these. Prosite does not have a nice API interface like UniProt, but the principles of using R's httr package to send POST requests and retrieve the results are the same. Getting data informally from Webpages is called screenscraping and really a life-saving skill. The first step to capture the data from this page via screenscraping is to look into the HTML code of the page.
(I am writing this section from the perspective of the Chrome browser - I don't think other browsers have all of the functionality that I am describing here. You may need to install Chrome to try this...)
- Use the menu and access View → Developer → View Source. Scroll through the page. You should easily be able to identify the data table. That's fair enough: each of the lines contain the UniProt ID and we should be able to identify them. But how to send the request to get this page in the first place?
- Use the browser's back button, and again: View → Developer → View Source. This is the page that accepts user input in a so called
formvia several different types of elements: "radio-buttons", a "text-box", "check-boxes", a "drop down menu" and a "submit" button. We need to figure out what each of the values are so that we can construct a validPOSTrequest. If we get them wrong, in the wrong order, or have parts missing, it is likely that the server will simply ignore our request. These elements are much harder to identify thean the lines of feature information, and it's really easy to get them wrong, miss something and get no output. But Chrome has a great tool to help us: it allows you to see the exact, assembledPOSTheader that it sent to the Prosite server!
- On the scanProsite page, open View → Developer → Developer Tools in the Chrome menu. Then click again on START THE SCAN. The Developer Tools page will show you information about what just happened in the transaction it negotiated to retrieve the results page. Click on the Network tab, and then on the top element:
PSScan.cgi. This contains the form data. Then click on the Headers tab and scroll down until you see the Request Payload. This has all the the requiredPOSTelements nicely spelled out. No guesswork required. What worked from the browser should work the same way from an R script. Analogous to our UniProt fetch code, we create aPOSTquery:
URL <- "http://prosite.expasy.org/cgi-bin/prosite/PSScan.cgi"
response <- POST(URL,
body = list(meta = "opt1",
meta1_protein = "opt1",
seq = "P39678",
skip = "on",
output = "tabular"))
# Note how the list-elements correspond to the page header's
# Request Payload. We include everything but the value of the
# submit button (which is for display only) in our POST
# request.
# Send off this request, and you should have a response in a few
# seconds.
# The text contents of the response is available with the
# content() function:
content(response, "text")
# ... should show you the same as the page contents that
# you have seen in the browser. Now we need to extract
# the data from the page: we need regular expressions, but
# only simple ones. First, we strsplit() the response into
# individual lines, since each of our data elements is on
# its own line. We simply split on the "\\n" newline character.
lines <- unlist(strsplit(content(response, "text"), "\\n"))
head(lines)
# Now we define a query pattern for the lines we want:
# we can use the uID, bracketed by two "|" pipe
# characters:
pattern <- paste("\\|", uID, "\\|", sep="")
# ... and select only the lines that match this
# pattern:
lines <- lines[grep(pattern, lines)]
lines
# ... captures the four lines of output.
# Now we break the lines apart into
# apart in tokens: this is another application of
# strsplit(), but this time we split either on
# "pipe" characters, "|" OR on tabs "\t". Look at the
# regex "\\t|\\|" in the strsplit() call:
strsplit(lines[1], "\\t|\\|")
# Its parts are (\\t)=tab (|)=or (\\|)=pipe.
# Both "t" and "|" need to be escaped with a backslash.
# "t" has to be escaped because we want to match a tab (\t),
# not the literal character "t". And "|" has to be escaped
# because we mean the literal pipe character, not its
# usual (special) meaning OR. Thus sometimes the backslash
# turns a special meaning off, and sometimes it turns a
# special meaning on. Unfortunately there's no easy way
# to tell - you just need to remember the characters - or
# have a reference handy. The special characters are
# (){}[]^$?*+.|&- ... and some of them have different
# meanings depending on where in the regex they are.
# Let's put the tokens into named slots of a vector.
features <- list()
for (line in lines) {
tokens <- unlist(strsplit(line, "\\t|\\|"))
features <- rbind(features, c(uID = tokens[2],
start = tokens[4],
end = tokens[5],
psID = tokens[6],
psName = tokens[7]))
}
featuresThis forms the base of a function that collects the features automatically from a PrositeScan result. We still need to do a bit more on the database part, but this is mostly bookkeeping:
- We need to put the feature annotations into a database table and link them to a protein ID and to a description of the feature itself.
- We need a function that extracts feature sequences in FASTA format.
- And, since we are changing the structure of the database, we need a way to migrate your old database contents to a newer version.
I don't think much new can be learned from this, so I have written those functions and put them into dbUtilities.R But you can certainly learn something from having a look at the code of
fetchPrositeFeatures()addFeatureToDB()getFeatureFASTA()
Also, have a quick look back at our database schema: this update has implemented the proteinFeature and the feature table. Do you remember what they were good for?
Time for a database update. You must be up to date with the latest version of dbUtilities.r for this to work. When you are, execute the following steps:
updateVerifiedFile("363ffbae3ff21ba80aa4fbf90dcc75164dbf10f8")
# Make a backup copy of your protein database.
# Load your protein database. Then merge the data in your database
# with the updated reference database. (Obviously, substitute the
# actual filename in the placeholder strings below. And don't type
# the angled brackets!)
<my-new-database> <- mergeDB(<my-old-database>, refDB)
# check that this has worked:
str(<my-new-database>)
# and save your database.
save(<my-new-database>, file="<my-DB-filename.02>.RData")
# Now, for each of your proteins, add the domain annotations to
# the database. You could write a loop to do this but it's probably
# better to check the results of each annotation before committing
# it to the database. So just paste the UniProt Ids as argument of
# the function fetchPrositeFeatures(), execute and repeat.
features <- fetchPrositeFeatures(<one-of-my-proteins-uniProt-IDs>)
refDB <- addFeatureToDB(refDB, features)
# When you are done, save your database.Finally, we can create a sequence selection of APSES domains
from our reference proteins. The function getFeatureFasta()
- accepts a feature name such as
"HTH_APSES; - finds the corresponding feature ID;
- finds all matching entries in the proteinFeature table;
- looks up the start and end position of each feature;
- fetches the corresponding substring from the sequence entries;
- adds a meaningful header line; and
- writes everything to output.
... so that you can simply execute:
cat(getFeatureFasta(<my-new-database>, "HTH_APSES"))Here are the first five sequences from that result:
>CC1G_01306_COPCI HTH_APSES 6:112
IFKATYSGIPVYEMMCKGVAVMRRRSDSWLNATQILKVAGFDKPQRTRVLEREVQKGEHE
KVQGGYGKYQGTWIPLERGMQLAKQYNCEHLLRPIIEFTPAAKSPPL
>CNBB4890_CRYNE HTH_APSES 17:123
IYKATYSGVPVYEMVCRDVAVMRRRSDAYLNATQILKVAGFDKPQRTRVLEREVQKGEHE
KVQGGYGKYQGTWIPIERGLALAKQYGVEDILRPIIDYVPTSVSPPP
>COCMIDRAFT_338_BIPOR HTH_APSES 9:115
IYSATYSNVPVYECNVNGHHVMRRRADDWINATHILKVADYDKPARTRILEREVQKGVHE
KVQGGYGKYQGTWIPLEEGRGLAERNGVLDKMRAIFDYVPGDRSPPP
>WALSEDRAFT_68476_WALME HTH_APSES 83:192
IYSAVYSGVGVYEAMIRGIAVMRRRADGYMNATQILKVAGVDKGRRTKILEREILAGLHE
KIQGGYGKYQGTWIPFERGRELALQYGCDHLLAPIFDFNPSVMQPSAGRS
>PGTG_08863_PUCGR HTH_APSES 90:196
IYKATYSGVPVLEMPCEGIAVMRRRSDSWLNATQILKVAGFDKPQRTRVLEREIQKGTHE
KIQGGYGKYQGTWVPLDRGIDLAKQYGVDHLLSALFNFQPSSNESPP
[...]
At the bottom of these sequences, you should see the APSES sequences from
YFO, in particular the Mbp1 RBM sequence from YFO. Email me if you have trouble getting to that stage.
We'll need to align these sequences with the template...
Template choice and template sequence
The SWISS-MODEL server provides several different options for constructing homology models. The easiest option requires only a target sequence as input. In this mode the program will automatically choose suitable templates and create an input alignment. I would argue however that that is not the best way to use such a service: template choice and alignment both may be significantly influenced by biochemical reasoning, and an automated algorithm cannot make the necessary decisions. Should you use a structure of reduced resolution that however has a ligand bound? Should you move an indel from an active site to a loop region even though the sequence similarity score might be less? Questions like that may yield answers that are different from the best choices an automated algorithm could make. But Swiss Model is flexible and allows us to upload an explicit alignment between target and template. Please note: the model you will produce is "easy" - the sequence similarity is high and there are no indels to consider, the automated mode would have done just as well. But the strategy we pursue here is suitable also for much more difficult problems. The automated strategy probably is not.
Template choice is the first step. Often more than one related structure can be found in the PDB. The degree of sequence identity is the most important criterion, but there are many other factors to consider. Please refer to the template choice principles page on this Wiki where I discuss more details and alternatives. To find related structures, you can search the PDB itself through its Advanced Search interface; for example one can search for sequence similarity with a BLAST search, or search for structural similarity by accessing structures according to their CATH or SCOP classification. But the BLAST search is probably the method of choice: after all, the most important measure of the probability of success for homology modelling is sequence similarity.
Defining a template means finding a PDB coordinate set that has sufficient sequence similarity to your target that you can build a model based on that template. To find suitable PDB structures, we will perform a BLAST search at the PDB.
Task:
- Retrieve your YFO's Mbp1 RBM APSES domain sequence from the FASTA selection you have just prepared. This YFO sequence is your target sequence.
- Navigate to the PDB.
- Click on Advanced to enter the advanced search interface.
- Open the menu to Choose a Query Type:
- Find the Sequence features section and choose Sequence (BLAST...)
- Paste your target sequence into the Sequence field, select not to mask low-complexity regions and Submit Query. Since the E-value is set rather high by default, you will get a number of low-confidence hits as well as the actual homologs, these have very low E-values.
All hits that are homologs are potentially suitable templates, but some are more suitable than others. Consider how the coordinate sets differ and which features would make each more or less suitable for creating a homology model: you should consider ...
- sequence similarity to your target
- size of expected model (= length of alignment)
- presence or absence of ligands
- experimental method and quality of the data set
Sequence similarity is the most important, but we can have the PDB tabulate the other features concisely for this task.
- There is a menu to create Reports: - select customizable table.
- Select (at least) the following information items:
- Structure Summary
- Experimental Method
- Sequence
- Chain Length
- Ligands
- Ligand Name
- Biological details
- Macromolecule Name
- refinement Details
- Resolution
- R Work
- R free
- click: Create report.
Unfortunately you don't get the E-values into the report, and those should strongly influence your final decision. However in our case the sequences and therefore the E-values of the top three hits are all the same. And there is a new structure from January 2015, with a lower resolution. Some of the sequences have a longer chain-length ... but those are only disordered residues (otherwise these would be better suited templates; regrettably, you'd need to check that in the real world, there is no automatic tool to evaluate disorder and its effects on template choice). In my opinion that leaves pretty much only one unambiguous choice for our template: 1BM8.
- Finally
- Click on the 1BM8 ID to navigate to the structure page for the template and save the FASTA sequence to your computer. This is the template sequence.
Sequence numbering
It is not straightforward at all how to number sequence in such a project. A "natural" numbering starts with the start-codon of the full length protein and goes sequentially from there. However, this does not map exactly to other numbering schemes we have encountered. As you know the first residue of the APSES domain (as defined by CDD) is not Residue 1 of the Mbp1 protein. The first residue of the 1BM8 FASTA file (one of the related PDB structures) is the fourth residue of the Mbp1 protein. The first residue in the structure is GLN 3, therefore Q is the first residue in a FASTA sequence derived from the cordinate section of the PDB file (the ATOM records. In the 1MB1 structure, the original N-terminal amino acids are present in the molecule, therefore they are present in the FASTA file which starts with MSNQIY..., but they are disordered in the structure and no coordinates are present for M and S. A sequence derived explicitly from the coordinates is therefore different from the reported FASTA sequence, which is really bad because that is what the modeling program has to work with ... and so on. It can get complicated. You need to remember: a sequence number is not absolute, but assigned in a particular context and you need to be careful how to do this.
Fortunately, the numbering for the residues in the coordinate section of our target structure corresponds not to its FASTA sequence, but to the numbering of the gene. Otherwise we would need to renumber the sequence (e.g. by using the bio3D R package). If we would not do this, the sequence numbers in the model might not correspond to the sequence numbers of our target.
The input alignment
The sequence alignment between target and template is the single most important factor that determines the quality of your model. No comparative modeling process will repair an incorrect alignment; it is useful to consider a homology model rather like a three-dimensional map of a sequence alignment rather than a structure in its own right. In a homology modeling project, typically the largest amount of time should be spent on preparing the best possible alignment. Even though automated servers like the SwissModel server will align sequences and select template structures for you, it would be unwise to use these just because they are convenient. You should take advantage of the much more sophisticated alignment methods available. Analysis of wrong models can't be expected to produce right results.
The best possible alignment is usually constructed from a multiple sequence alignment that includes at least the target and template sequence and other related sequences as well. The additional sequences are an important aid in identifying the correct placement of insertions and deletions. Your alignment should have been carefully reviewed by you and wherever required, manually adjusted to move insertions or deletions between target and template out of the secondary structure elements of the template structure.
In most of the Mbp1 orthologues, we do not observe indels in the APSES domain regions. Evolutionary pressure on the APSES domains has selected against indels in the more than 600 million years these sequences have evolved independently in their respective species. To obtain an alignment between the template sequence and the target sequence from your species, proceed as follows.
Task:
Choose one of the following options to align your target and template sequence. Make sure your template sequence is included, i.e. the FASTA sequence of 1BM8.
- In Jalview...
- Load your APSES domain sequences plus the 1BM8 sequence in Jalview. Include the sequence of your template protein and align using Muscle.
- Delete all sequence you no longer need, i.e. keep only the APSES domains of the target (from your species) and the template (from the PDB) and choose Edit → Remove empty columns. This is your input alignment.
- Choose File→Output to textbox→FASTA to obtain the aligned sequences. They should both have exactly the same length, i.e. N- or C- termini have to be padded by hyphens if the original sequences had different length. Save the sequences in a text-file.
- Using a different MSA program
- Copy the FASTA formatted sequences of the Mbp1 proteins in the reference species from the Reference APSES domain page.
- Access the MSA tools page at the EBI.
- Paste the Mbp1 sequence set, your target sequence and the template sequence into the input form.
- Run an alignment (I like T-coffee) and save the output.
- Using the R bioconductor MSA package that you used previously.
Refer back to the page if you are lacking notes how to go about this.
Whatever method you use: the result should be a two sequence alignment in multi-FASTA format, that was constructed from a number of supporting sequences and that contains your aligned target and template sequence. This is your input alignment for the homology modeling server. For a Schizosaccharomyces pombe model, which I am using as an example here, it looks like this:
>1BM8_A QIYSARYSGVDVYEFIHSTGSIMKRKKDDWVNATHILKAANFAKAKRTRI LEKEVLKETHEKVQGGFGKYQGTWVPLNIAKQLAEKFSVYDQLKPLFDF >Mbp1_SCHPO 2-100 NP_593032 AVHVAVYSGVEVYECFIKGVSVMRRRRDSWLNATQILKVADFDKPQRTRV LERQVQIGAHEKVQGGYGKYQGTWVPFQRGVDLATKYKVDGIMSPILSL
.
- Note that you can enter the lowest acceptable match % separately for query and target. This means: what percentage of secondary structure elements would need to be matched in either query or target to produce a hit. Keep that value at 80 for our query, since we would want to find structures with almost all of the elements of the winged helix motif. Set the match to 10 % for the target, since we are interested in such domains even if they happen to be small subdomains of large proteins.
- Keep the Precision at normal. Precision and % query match could be relaxed if we wanted to find more structures.
- Finally click on: Submit your query.
- On the results page, click on the index number (in the left-hand column) of the top hit that is not one of our familiar Mbp1 structures to get a detailed view of the result. Most likely this is
1wq2:a, an enzyme. Click on View Superposed. This will open a window with the structure coordinates superimposed in the Jmol molecular viewer. Control-click anywhere in the window area to open a menu of viewing options. Select Style → Stereographic → Wall-eyed viewing. Select Trace as the rendering. Then study the superposition. You will note that the secondary structure elements match quite well, but does this mean we have a DNA-binding domain in this sulfite reductase?
}}
All in all this appears to be well engineered software! It gives you many options to access result details for further processing. I think this can be put to very good use. But for our problem, we would have to search through too many structures because, once again, we can't tell which ones of the hits are DNA binding domains, especially domains for which the structure of a complex has been solved.
APSES domains represent one branch of the tree of helix-turn-helix (HTH) DNA binding modules. (A review on HTH proteins is linked from the resources section at the bottom of this page). Winged Helix domains typically bind their cognate DNA with a "recognition helix" which precedes the beta hairpin and binds into the major groove; additional stabilizing interactions are provided by the edge of a beta-strand binding into the minor groove. This is good news: once we have determined that the APSES domain is actually an example of a larger group of transcription factors, we can compare our model to a structure of a protein-DNA complex. Superfamilies of such structural domains are compiled in the CATH database. Unfortunately CATH itself does not provide information about whether the structures have been determined as complexes. But we can search the PDB with CATH codes and restrict the results to complexes. Essentially, this should give us a list of all winged helix domains for which the structure of complexes with DNA have been determined. This works as follows:
Task:
- For reference, access CATH domain superfamily 1.10.10.10; this is the CATH classification code we will use to find protein-DNA complexes. Click on Superfamily Superposition to get a sense of the structural core of the winged helix domain.
- Navigate to the PDB home page and follow the link to Advanced Search
- In the options menu for Choose a Query Type select Structure Features → CATH Classification Browser. A window will open that allows you to navigate down through the CATH tree. You can view the Class/Architecture/Topology names on the CATH page linked above. Click on the triangle icons (not the text) for Mainly Alpha → Orthogonal Bundle → ARC repressor mutant, subunit A then click on the link to winged helix repressor DNA binding domain. Or, just enter "winged helix" into the search field. This subquery should match more than 550 coordinate entries.
- Click on the (+) button behind Add search criteria to add an additional query. Select the option Structure Features → Macromolecule type. In the option menus that pop up, select Contains Protein→Yes, Contains DNA→Yes, Contains RNA→Ignore, Contains DNA/RNA hybrid→Ignore. This selects files that contain Protein-DNA complexes.
- Check the box below this subquery to Remove Similar Sequences at 90% identity and click on Submit Query. This query should retrieve more than 100 complexes.
- Scroll down to the beginning of the list of PDB codes and locate the Reports menu. Under the heading View select Gallery. This is a fast way to obtain an overview of the structures that have been returned. Adjust the number of Results to see all 100 images and choose Options→Resize medium.
- Finally we have a set of winged-helix domain/DNA complexes, for comparison. Scroll through the gallery and study how the protein binds DNA.
First of all you may notice that in fact not all of the structures are really different, despite having requested only to retrieve dissimilar sequences, and not all images show DNA. This appears to be a deficiency of the algorithm. But you can also easily recognize how in most of the the structures the recognition helix inserts into the major groove of B-DNA (eg. 1BC8, 1CF7) and the wing - if clearly visible at all in the image - appears to make accessory interactions with the DNA backbone.. There is one exception: the structure 1DP7 shows how the human RFX1 protein binds DNA in a non-canonical way, through the beta-strands of the "wing". This is interesting since it suggests there is more than one way for winged helix domains to bind to DNA. We can therefore use structural superposition of your homology model and two of the winged-helix proteins to decide whether the canonical or the non-canonical mode of DNA binding seems to be more plausible for Mbp1 orthologues.
Preparation and superposition of a canonical complex
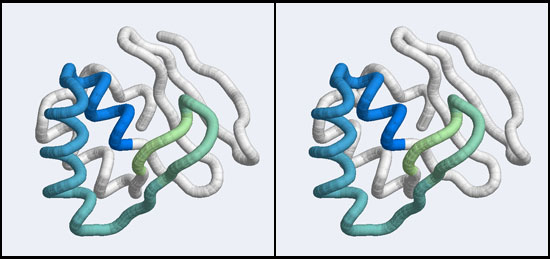
The structure we shall use as a reference for the canonical binding mode is the Elk-1 transcription factor.
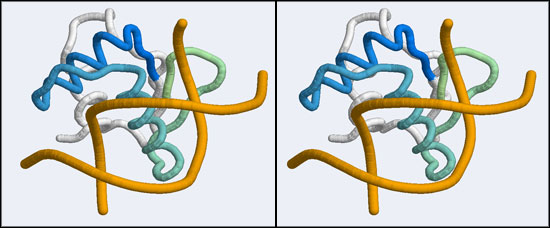
The 1DUX coordinate-file contains two protein domains and two B-DNA dimers in one asymmetric unit. For simplicity, you should delete the second copy of the complex from the PDB file. (Remember that PDB files are simply text files that can be edited.)
Task:
- Find the 1DUX structure in the image gallery and open the 1DUX structure explorer page in a separate window. Download the coordinates to your computer.
- Open the coordinate file in a text-editor (TextEdit or Notepad - NOT MS-Word!) and delete the coordinates for chains
D,EandF; you may also delete allHETATMrecords and theMASTERrecord. Save the file with a different name, e.g. 1DUX_monomer.pdb . - Open VMD and load your homology model. Turn off the axes, display the model as a Tube representation in stereo, and color it by Index. Then load your edited 1DUX file, display this coordinate set in a tube representation as well, and color it by ColorID in some color you like. It is important that you can distinguish easily which structure is which.
- You could use the Extensions→Analysis→RMSD calculator interface to superimpose the two strutcures IF you would know which residues correspond to each other. Sometimes it is useful to do exactly that: define exact correspondences between residue pairs and superimpose according to these selected pairs. For our purpose it is much simpler to use the Multiseq tool (and the structures are simple and small enough that the STAMP algorithm for structural alignment can define corresponding residue pairs automatically). Open the multiseq extension window, select the check-boxes next to both protein structures, and open the Tools→Stamp Structural Alignment interface.
- In the "'Stamp Alignment Options'" window, check the radio-button for Align the following ... Marked Structures and click on OK.
- In the Graphical Representations window, double-click on all "NewCartoon" representations for both molecules, to undisplay them.
- You should now see a superimposed tube model of your homology model and the 1DUX protein-DNA complex. You can explore it, display side-chains etc. and study some of the details of how a transcription factor recognizes and binds to its cognate DNA sequence. However, remember that your model's side-chain orientations have not been determined experimentally but inferred from the template, and that the template's structure was determined in the absence of bound DNA ligand.
- Orient and scale your superimposed structures so that their structural similarity is apparent, and the recognition helix can be clearly seen inserting into the DNA major groove. You may want to keep a copy of the image for future reference. Consider which parts of the structure appear to superimpose best. Note whether it is plausible that your model could bind a B-DNA double-helix in this orientation.
Preparation and superposition of a non-canonical complex
The structure displaying a non-canonical complex between a winged-helix domain and its cognate DNA binding site is the human Regulatory Factor X.
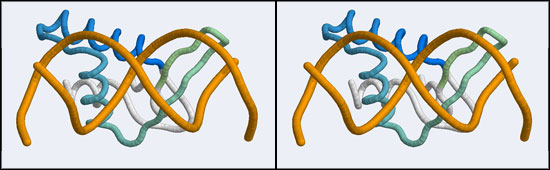
Before we can work with this however, we have to fix an annoying problem. If you download and view the 1DP7 structure in VMD, you will notice that there is only a single strand of DNA! Where is the second strand of the double helix? It is not in the coordinate file, because it happens to be exactly equivalent to the frist starnd, rotated around a two-fold axis of symmetry in the crystal lattice. We need to download and work with the so-called Biological Assembly instead. But there is a problem related to the way the PDB stores replicates in biological assemblies. The PDB generates the additional chains as copies of the original and delineates them with MODEL and ENDMDL records, just like in a multi-structure NMR file. The chain IDs and the atom numbers are the same as the original. The PDB file thus contains the same molecule in two different orientations, not two independent molecules. This is an important difference regarding how such molecules are displayed by VMD. If you try to use the biological unit file of the PDB, VMD does not recognize that there is a second molecule present and displays only one chain. And that looks exactly like the one we have seen before. We have to edit the file, extract the second DNA molecule, change its chain ID and then append it to the original 1DP7 structure[1]...
Task:
- On the structure explorer page for 1DP7, select the option Download Files → PDB File.
- Also select the option Download Files → Biological Assembly.
- Uncompress the biological assembly file.
- Open the file in a text editor.
- Delete everything except the second DNA molecule. This comes after the
MODEL 2line and has chain ID D. Keep theTERandENDlines. Save this with a new filename (e.g.1DP7_DNAonly.pdb). - Also delete all
HETATMrecords forHOH,PEGandEDO, as well as the entire second protein chain and theMASTERrecord. The resulting file should only contain the DNA chain and its copy and one protein chain. Save the file with a new name, eg.1DP7_BDNA.PDB. - Use a similar procedure as BIO_Assignment_Week_8#R code: renumbering the model in the last assignment to change the chain ID.
PDBin <- "1DP7_DNAonly.pdb"
PDBout <- "1DP7_DNAnewChain.pdb"
pdb <- read.pdb(PDBin)
pdb$atom[,"chain"] <- "E"
write.pdb(pdb=pdb,file=PDBout)- Use your text-editor to open both the
1DP7.pdbstructure file and the1DP7_DNAnewChain.pdb. Copy the DNA coordinates, paste them into the original file before theENDline and save. - Open the edited coordinate file with VMD. You should see one protein chain and a B-DNA double helix. (Actually, the BDNA helix has a gap, because the R-library did not read the BRDU nucleotide as DNA). Switch to stereo viewing and spend some time to see how amazingly beautiful the complementarity between the protein and the DNA helix is (you might want to display protein and nucleic in separate representations and color the DNA chain by Position → Radial for clarity) ... in particular, appreciate how not all positively charged side chains contact the phosphate backbone, but some pnetrate into the helix and make detailed interactions with the nucleobases!
- Then clear all molecules
- In VMD, open Extensions→Analysis→MultiSeq. When you run MultiSeq for the first time, you will be asked for a directory in which to store metadata. You can use the default, or a directory of your choice; you may subsequently skip all steps that ask you to install "required" databases locally since we will not need them for this task.
- Choose File→Import Data, browse to your directory and load one by one:
- -Your model;
- -The 1DUX complex;
- -The 1DP7 complex.
- Mark all three protein chains by selecting the checkbox next to their name and choose Tools→ STAMP structural alignment.
- Align the Marked Structures, choose a scanscore of 2 and scanslide of 5. Also choose Slow scan. You may have to play around with the setting to get the molecules to superimpose: but the can be superimposed quite well - at least the DNA-binding helices and the wings should line up.
- In the graphical representations window, double-click on the cartoon representations that multiseq has generated to undisplay them, also undisplay the Tube representation of 1DUX. Then create a Tube representation for 1DP7, and select a Color by ColorID (a different color that you like). The resulting scene should look similar to the one you have created above, only with 1DP7 in place of 1DUX and colored differently.
- Orient and scale your superimposed structures so that their structural similarity is apparent, and the differences in binding elements is clear. Perhaps visualizing a solvent accessible surface of the DNA will help understand the spatial requirements of the complex formation. You may want to keep a copy of the image for future reference. Note whether it is plausible that your model could bind a B-DNA double-helix in the "alternative" conformation.
-->
Links and resources
| Altenhoff & Dessimoz (2012) Inferring orthology and paralogy. Methods Mol Biol 855:259-79. (pmid: 22407712) |
|
[ PubMed ] [ DOI ] The distinction between orthologs and paralogs, genes that started diverging by speciation versus duplication, is relevant in a wide range of contexts, most notably phylogenetic tree inference and protein function annotation. In this chapter, we provide an overview of the methods used to infer orthology and paralogy. We survey both graph-based approaches (and their various grouping strategies) and tree-based approaches, which solve the more general problem of gene/species tree reconciliation. We discuss conceptual differences among the various orthology inference methods and databases, and examine the difficult issue of verifying and benchmarking orthology predictions. Finally, we review typical applications of orthologous genes, groups, and reconciled trees and conclude with thoughts on future methodological developments. |
- PDB file format (see the Coordinate Section if you are unsure about chain identifiers)
- Wikipedia on Structural Superposition (although the article is called "Structural Alignment")
- Reference sequences
Footnotes and references
- ↑ My apologies if this is tedious. But in the real world, we encounter such problems a lot and I would be remiss not to use this opportunity to let you practice how to fix the issue that could otherwise be a roadblock in a project of yours.
Ask, if things don't work for you!
- If anything about the assignment is not clear to you, please ask on the mailing list. You can be certain that others will have had similar problems. Success comes from joining the conversation.
- Do consider how to ask your questions so that a meaningful answer is possible:
- How to create a Minimal, Complete, and Verifiable example on stackoverflow and ...
- How to make a great R reproducible example are required reading.
| < Assignment 6 | Assignment 8 > |