WWW Reactome
Reactome |
URL
Abstract
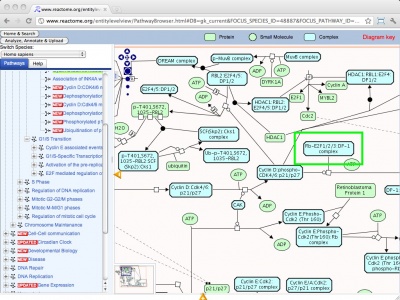
Reactome is a multi-site collaboration to develop an open source, curated bioinformatics database of human pathways and reactions. It includes annotations, pathways and tools for pathway browsing and analysis, including pathway assignment and overrepresentation analysis of user-supplied data sets. Making use of orthology prediction, Reactome also provides cross-species pathway inference for a large number of model organisms. The URL accesses the E2F mediated regulation of DNA replication.
Reference
| Croft et al. (2011) Reactome: a database of reactions, pathways and biological processes. Nucleic Acids Res 39:D691-7. (pmid: 21067998) |
|
[ PubMed ] [ DOI ] Reactome (http://www.reactome.org) is a collaboration among groups at the Ontario Institute for Cancer Research, Cold Spring Harbor Laboratory, New York University School of Medicine and The European Bioinformatics Institute, to develop an open source curated bioinformatics database of human pathways and reactions. Recently, we developed a new web site with improved tools for pathway browsing and data analysis. The Pathway Browser is an Systems Biology Graphical Notation (SBGN)-based visualization system that supports zooming, scrolling and event highlighting. It exploits PSIQUIC web services to overlay our curated pathways with molecular interaction data from the Reactome Functional Interaction Network and external interaction databases such as IntAct, BioGRID, ChEMBL, iRefIndex, MINT and STRING. Our Pathway and Expression Analysis tools enable ID mapping, pathway assignment and overrepresentation analysis of user-supplied data sets. To support pathway annotation and analysis in other species, we continue to make orthology-based inferences of pathways in non-human species, applying Ensembl Compara to identify orthologs of curated human proteins in each of 20 other species. The resulting inferred pathway sets can be browsed and analyzed with our Species Comparison tool. Collaborations are also underway to create manually curated data sets on the Reactome framework for chicken, Drosophila and rice. |