BIO Assignment Week 3
Assignment for Week 3
Sequence Analysis
| < Assignment 2 | Assignment 4 > |
Note! This assignment is currently active. All significant changes will be announced on the mailing list.
Concepts and activities (and reading, if applicable) for this assignment will be topics on next week's quiz.
Contents
Introduction
Store
The Protein datamodel
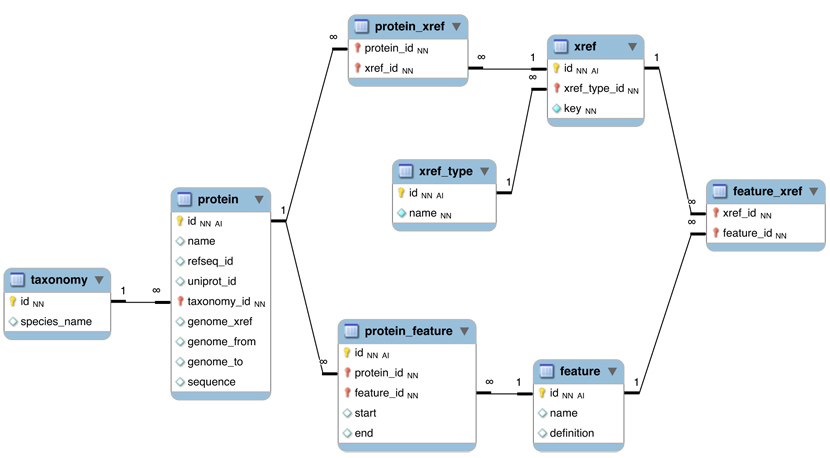
Here is a revised schema for the initial model that we developed in class:
- The
proteintable is at the centre. Its primary key is a unique integer. We store the NCBI RefSeq ID and the Uniprot ID in the table. We would not call them "foreign keys", since the information they reference is not in our schema but at the NCBI/EBI. Thegenome_xreffield will hold the ID for the actual genomic sequence, where the coding region of the gene is identified by its start ("from") and end ("to"). - The
taxonomytable holds information about species. We use the NCBI taxonomy ID as its primary key. The same key appears in the protein table as the foreign key that links the protein with its proper taxonomy information. - The
protein_featuretable links a protein with all the features that we annotate it with. Start and end coordinates identify the region of the sequence we have annotated. In principle these attributes could be NULL. - The
featuretable contains a feature name and a textual definition. Again, these attributes could theoretically be NULL. - The
protein_xreftable and thefeature_xreftable connect cross-reference data. - The
xreftable needs to hold two attributes: the type of cross-reference (e.g. PubMed), from which we would construct an URL or other accession method, and the actual key (e.g.10747782). The type of cross-reference needs some thought however: this field needs a "controlled vocabulary" to ensure that the same cross-reference is described consistently with the same string (pubmed? PubMed? Pub-Med?). There are a number of ways to achieve this, the best way is to store these types in their own table -xref_type- and link to that table via a foreign key.
Implementing the Data Model in R
To actually implement the model in R we will create the tables as data frames, and we will collect them in a list. We don't have to keep the tables in a list - we could also keep them as independent objects, but with a table named "protein" we should be worried of inadvertently overwriting the table. A list is neater, more flexible (we might want to have several of these), it reflects our intent about the model better, and doesn't require very much more typing. For now, to keep things simple, we will implement only two tables: protein and taxonomy. We'll add the rest when we actually need them.
db <- list() # we'll call our database "db" and start with an empty list
db$protein <- data.frame(id = 1,
name = "Mbp1",
refseq_id = "NP_010227",
uniprot_id = "P39678",
taxonomy_id = 4932,
genome_xref = "NC_001136.10",
genome_from = 352877,
genome_to = 355378,
sequence = "...",
stringsAsFactors = FALSE)
db$taxonomy <- data.frame(id = 4932,
species_name = "Saccharomyces cerevisiae",
stringsAsFactors = FALSE)
# Let's add a second protein:
db$protein <- rbind(db$protein,
data.frame(id = 2,
name = "Res2",
refseq_id = "NP_593032",
uniprot_id = "P41412",
taxonomy_id = 4896,
genome_xref = "NC_003424.3",
genome_from = 686543,
genome_to = 689179,
sequence = "...",
stringsAsFactors = FALSE))
db$taxonomy <- rbind(db$taxonomy,
data.frame(id = 4896,
species_name = "Schizosaccharomyces pombe",
stringsAsFactors = FALSE))
str(db)
# you can look at the contents of the tables in the usual way we would access
# elements from lists and dataframes. Here are some examples:
db$protein
db$protein$refseq_id
db$protein[,"name"]
db$taxonomy
db$taxonomy[1,]
# Here is an example to look up information in one table,
# based on a condition in another table:
# what is the species name for the protein
# whose name is "Mbp1"?
# First, get the taxonomy_id for the Mbp1 protein. This is
# the key we need for the taxonomy table. We find it in a cell in the
# table: db$protein[<row>, <column>]
# <row> is that row for which the value in
# the "name" column is Mbp1:
db$protein$name == "Mbp1" # TRUE FALSE
# The <column> is called "taxonomy_id". Simply
# insert these two expressions in the square
# brackets.
db$protein[db$protein$name == "Mbp1", "taxonomy_id"]
# Assign the taxonomy_id value ...
x <- db$protein[db$protein$name == "Mbp1", "taxonomy_id"]
# ... and fetch the species_name value from db$taxonomy
db$taxonomy[db$taxonomy$id == x, "species_name"]As you see we can edit any component of our data model directly by assigning new values to the element. But in general that's a really bad idea, since it is far too easy to bring the whole model into an inconsistent state. It's much better to write functions that get and set data elements, not only to keep the data consistent, but also because it is much easier, if the model ever changes, to simply edit the function code, rather than having to repeat every single data entry by hand.
What would an setData function have to look like? It should
- create a new entry if the requested row of a table does not exist yet;
- update data if the protein exists;
- perform consistency checks (i.e. check that the data has the correct type);
- perform sanity checks (i.e. check that data values fall into the expected range);
- perform completeness checks (i.e. handle incomplete data)
Let's start simple, and create a set- function to add new values to existing sequence data. Also, for clarity, we'll forgo many checks. The first thing we should do is to add the actual sequence.
We only entered a placeholder for the sequence field. Sequences come in many different flavours when we copy them from a Webpage: there can be whitespace, carriage returns, numbers, they can be upper-case, lower-case mixed-case ...
What we want in our sequence data field is one string that contains the entire sequence, and nothing but upper-case, amino-acid code letters.
We'll need to look at how we work with strings in R, how we identify and work with patterns in strings. This is a great time to introduce regular expressions.
A Brief First Encounter of Regular Expressions
- Regular expressions are a concise description language to define patterns for pattern-matching in strings.
Truth be told, many programmers have a love-hate relationship with regular expressions. The syntax of regular expressions is very powerful and expressive, but also terse, not always intuitive, and sometimes hard to understand. I'll introduce you to a few principles here that are quite straightforward and they will probably cover 99% of the cases you will encounter.
Here is our test-case: the sequence of Mbp1, copied from the Genbank Webpage.
1 msnqiysary sgvdvyefih stgsimkrkk ddwvnathil kaanfakakr trilekevlk
61 ethekvqggf gkyqgtwvpl niakqlaekf svydqlkplf dftqtdgsas pppapkhhha
121 skvdrkkair sastsaimet krnnkkaeen qfqsskilgn ptaaprkrgr pvgstrgsrr
181 klgvnlqrsq sdmgfprpai pnssisttql psirstmgpq sptlgileee rhdsrqqqpq
241 qnnsaqfkei dledglssdv epsqqlqqvf nqntgfvpqq qssliqtqqt esmatsvsss
301 pslptspgdf adsnpfeerf pgggtspiis miprypvtsr pqtsdindkv nkylsklvdy
361 fisnemksnk slpqvllhpp phsapyidap idpelhtafh wacsmgnlpi aealyeagts
421 irstnsqgqt plmrsslfhn sytrrtfpri fqllhetvfd idsqsqtvih hivkrksttp
481 savyyldvvl skikdfspqy rielllntqd kngdtalhia skngdvvffn tlvkmgaltt
541 isnkegltan eimnqqyeqm miqngtnqhv nssntdlnih vntnnietkn dvnsmvimsp
601 vspsdyityp sqiatnisrn ipnvvnsmkq masiyndlhe qhdneikslq ktlksisktk
661 iqvslktlev lkesskdeng eaqtnddfei lsrlqeqntk klrkrliryk rlikqkleyr
721 qtvllnklie detqattnnt vekdnntler lelaqeltml qlqrknklss lvkkfednak
781 ihkyrriire gtemnieevd ssldvilqtl iannnknkga eqiitisnan sha
//
Task:
Navigate to http://regexpal.com and paste the sequence into the lower box. This site is one of a number of online regular expression testers; their immediate, visual feedback is invaluable when you are developing regular expression patterns.
Lets try some expressions:
- Most characters are matched literally.
- Type "
a" in to the upper box and you will see all "a" characters matched. Then replaceawithq. - Now type "
aa" instead. Thenkrnnkk. Sequences of characters are also matched literally.
- We can specify more than one character to match if we place it in square brackets.
[lq]matcheslORq.[familyvw]matches hydrophobic amino acids.
- Within square brackets, we can specify "ranges".
[1-5]matches digits from 1 to 5.
- Within square brackets, we can specify characters that should NOT be matched, with the caret,
^. [^0-9]matches everything EXCEPT digits.[^a-z]matches everything that is not a lower-case letter. That's what we need (try it).
One of the R functions that uses regular expressions is the function gsub(). It replaces characters that match a "regex" with other characters. That is useful for our purpose: we can
- match all characters that are NOT a letter, and
- replace them by - nothing: the empty string
"".
This deletes them.
Task:
Try this code:
seq <- " # copy/paste from GenBank
1 msnqiysary sgvdvyefih stgsimkrkk ddwvnathil kaanfakakr trilekevlk
61 ethekvqggf gkyqgtwvpl niakqlaekf svydqlkplf dftqtdgsas pppapkhhha
121 skvdrkkair sastsaimet krnnkkaeen qfqsskilgn ptaaprkrgr pvgstrgsrr
181 klgvnlqrsq sdmgfprpai pnssisttql psirstmgpq sptlgileee rhdsrqqqpq
241 qnnsaqfkei dledglssdv epsqqlqqvf nqntgfvpqq qssliqtqqt esmatsvsss
301 pslptspgdf adsnpfeerf pgggtspiis miprypvtsr pqtsdindkv nkylsklvdy
361 fisnemksnk slpqvllhpp phsapyidap idpelhtafh wacsmgnlpi aealyeagts
421 irstnsqgqt plmrsslfhn sytrrtfpri fqllhetvfd idsqsqtvih hivkrksttp
481 savyyldvvl skikdfspqy rielllntqd kngdtalhia skngdvvffn tlvkmgaltt
541 isnkegltan eimnqqyeqm miqngtnqhv nssntdlnih vntnnietkn dvnsmvimsp
601 vspsdyityp sqiatnisrn ipnvvnsmkq masiyndlhe qhdneikslq ktlksisktk
661 iqvslktlev lkesskdeng eaqtnddfei lsrlqeqntk klrkrliryk rlikqkleyr
721 qtvllnklie detqattnnt vekdnntler lelaqeltml qlqrknklss lvkkfednak
781 ihkyrriire gtemnieevd ssldvilqtl iannnknkga eqiitisnan sha
//
"
seq # "\n" means: line-break
seq <- gsub("[^a-zA-Z]", "", seq) #replace all non-letters with ""
seq
# Now to change the sequence to upper-case. R has toupper()
# and tolower().
toupper(seq)
# Lets put this in a function:
sanitizeSequence <- function(s) {
return(toupper(gsub("[^a-zA-Z]", "", s)))
}
# try it:
test <- "f123 a$^&&*)m. i l马马虎虎 é yßv+w"
sanitizeSequence(test)A Function to Set Values
Building on this function, we can write a function to set values in our protein table.
setDB <- function(database,
table,
id,
name = "",
refseq_id = "",
uniprot_id = "",
taxonomy_id = "",
genome_xref = "",
genome_from = "",
genome_to = "",
sequence = "",
species_name = "") {
if (missing(database) | missing(table) | missing(id)) {
stop("database, table and id have to be specified")
}
if (table == "protein") {
row <- which(database$protein$id == id)
if (name != "") { database$protein[row, "name"] <- name }
if (refseq_id != "") { database$protein[row, "refseq_id"] <- refseq_id}
if (uniprot_id != "") { database$protein[row, "uniprot_id"] <- uniprot_id}
if (taxonomy_id != "") { database$protein[row, "taxonomy_id"] <- taxonomy_id}
if (genome_xref != "") { database$protein[row, "genome_xref"] <- genome_xref}
if (genome_from != "") { database$protein[row, "genome_from"] <- genome_from}
if (genome_to != "") { database$protein[row, "genome_to"] <- genome_to}
if (sequence != "") { database$protein[row, "sequence"] <- sanitizeSequence(sequence)}
}
if (table == "taxonomy") {
row <- which(database$taxonomy$id == id)
if (species_name != "") { database$taxonomy[row, "species_name"] <- species_name }
}
return(database)
}
# Note that this code could be significantly improved: we are
# not doing any tests that the values are sane.
# But for now I just want to keep the code simple.
db$protein[1,]
db <- setDB(database = db, table="protein", id=1, name="Armadillo")
db$protein[1,]
db <- setDB(database = db, table="protein", id=1, name="Mbp1")
# With this, we can update our entries to set the sequence to its actual value.
seq
seq <- "
1 msnqiysary sgvdvyefih stgsimkrkk ddwvnathil kaanfakakr trilekevlk
61 ethekvqggf gkyqgtwvpl niakqlaekf svydqlkplf dftqtdgsas pppapkhhha
121 skvdrkkair sastsaimet krnnkkaeen qfqsskilgn ptaaprkrgr pvgstrgsrr
181 klgvnlqrsq sdmgfprpai pnssisttql psirstmgpq sptlgileee rhdsrqqqpq
241 qnnsaqfkei dledglssdv epsqqlqqvf nqntgfvpqq qssliqtqqt esmatsvsss
301 pslptspgdf adsnpfeerf pgggtspiis miprypvtsr pqtsdindkv nkylsklvdy
361 fisnemksnk slpqvllhpp phsapyidap idpelhtafh wacsmgnlpi aealyeagts
421 irstnsqgqt plmrsslfhn sytrrtfpri fqllhetvfd idsqsqtvih hivkrksttp
481 savyyldvvl skikdfspqy rielllntqd kngdtalhia skngdvvffn tlvkmgaltt
541 isnkegltan eimnqqyeqm miqngtnqhv nssntdlnih vntnnietkn dvnsmvimsp
601 vspsdyityp sqiatnisrn ipnvvnsmkq masiyndlhe qhdneikslq ktlksisktk
661 iqvslktlev lkesskdeng eaqtnddfei lsrlqeqntk klrkrliryk rlikqkleyr
721 qtvllnklie detqattnnt vekdnntler lelaqeltml qlqrknklss lvkkfednak
781 ihkyrriire gtemnieevd ssldvilqtl iannnknkga eqiitisnan sha
//
"
db <- setDB(database = db, table="protein", id=1, sequence=seq)
seq <- "
1 maprssavhv avysgvevye cfikgvsvmr rrrdswlnat qilkvadfdk pqrtrvlerq
61 vqigahekvq ggygkyqgtw vpfqrgvdla tkykvdgims pilsldideg kaiapkkkqt
121 kqkkpsvrgr rgrkpsslss stlhsvnekq pnssisptie ssmnkvnlpg aeeqvsatpl
181 paspnallsp ndntikpvee lgmleapldk yeeslldffl hpeegripsf lyspppdfqv
241 nsvidddght slhwacsmgh iemiklllra nadigvcnrl sqtplmrsvi ftnnydcqtf
301 gqvlellqst iyavdtngqs ifhhivqsts tpskvaaaky yldcilekli siqpfenvvr
361 lvnlqdsngd tslliaarng amdcvnslls ynanpsipnr qrrtaseyll eadkkphsll
421 qsnsnashsa fsfsgispai ispscsshaf vkaipsissk fsqlaeeyes qlrekeedli
481 ranrlkqdtl neisrtyqel tflqknnpty sqsmenlire aqetyqqlsk rlliwlearq
541 ifdlerslkp htslsisfps dflkkedgls lnndfkkpac nnvtnsdeye qlinkltslq
601 asrkkdtlyi rklyeelgid dtvnsyrrli amscginped lsleildave ealtrek
"
db <- setDB(database = db, table="protein", id=2, sequence=seq)
A Function to Add New Entries
The function to add a new entry to the database looks quite similar. But we have to add a test whether the species name has been entered when we add a species with a new taxonomy ID.
addToDB <- function(database,
name = "",
refseq_id = "",
uniprot_id = "",
taxonomy_id = "",
genome_xref = "",
genome_from = "",
genome_to = "",
sequence = "",
species_name = "") {
if (missing(database) | missing(taxonomy_id)) {
stop("database and taxonomy_id have to be specified")
}
# handle taxonomy
if (! any(database$taxonomy$id == taxonomy_id)) { # new taxonomy_id
if (missing(species_name)) {
stop("taxonomy_id is new: species_name has to be specified")
}
else {
# add this species to the taxonomy table
database$taxonomy <- rbind(database$taxonomy,
data.frame(id = taxonomy_id,
species_name = species_name,
stringsAsFactors = FALSE))
}
}
# handle protein
# get a new protein id
pid <- max(database$protein$id) + 1
database$protein <- rbind(database$protein,
data.frame(id = pid,
name = name,
refseq_id = refseq_id,
uniprot_id = uniprot_id,
taxonomy_id = taxonomy_id,
genome_xref = genome_xref,
genome_from = genome_from,
genome_to = genome_to,
sequence = sanitizeSequence(sequence),
stringsAsFactors = FALSE))
return(database)
}
# Test this.
newSeq <- "
1 msgdktifka tysgvpvyec iinnvavmrr rsddwlnatq ilkvvgldkp qrtrvlerei
61 qkgihekvqg gygkyqgtwi pldvaielae ryniqgllqp itsyvpsaad spppapkhti
121 stsnrskkii padpgalgrs rratsietes evigaapnnv segsmspsps dissssrtps
181 plpadrahpl hanhalagyn grdannhary adiildyfvt enttvpslli npppdfnpdm
241 sidddehtal hwacamgrir vvklllsaga difrvnsnqq talmratmfs nnydlrkfpe
301 lfellhrsil nidrndrtvf hhvvdlalsr gkphaaryym etminrlady gdqladilnf
361 qddegetplt maararskrl vrlllehgad pkirnkegkn aedyiieder frsspsrtgp
421 agielgadgl pvlptsslht seagqrtagr avtlmsnllh sladsydsei ntaekkltqa
481 hgllkqiqte iedsakvaea lhheaqgvde erkrvdslql alkhainkra rddlerrwse
541 gkqaikrarl qaglepgals tsnatnapat gdqkskddak sliealpagt nvktaiaelr
601 kqlsqvqank telvdkfvar areqgtgrtm aayrrliaag cggiapdevd avvgvlcell
661 qeshtgarag aggerddrar dvammlkgag aaalaanaga p
"
db2 <- addToDB(db,
name = "UMAG_1122",
refseq_id = "XP_011392621",
uniprot_id = "A0A0D1DP35",
taxonomy_id = 5270,
genome_xref = "NC_026499.1",
genome_from = 150555,
genome_to = 152739,
sequence = newSeq,
species_name = "Ustilago maydis")As you can see, adding a new protein to our database is now rather compact.
Add Your YFO Sequence
Task:
In the last assignment, you discovered a protein via BLAST that is related to yeast Mbp1 in YFO. (If you haven't noted its ID in your journal, you'll have to redo the BLAST search.)
Add that protein to the database. Ask on the mailing list if you don't know how (but be specific) or if you don't know how to find particular information items.
Check that the protein has been properly added to all columns.
Analyze
Let us perform a few simple sequence analyses using the online EMBOSS tools. EMBOSS (the European Molecular Biology laboratory Open Software Suite) combines a large number of simple but fundamental sequence analysis tools. The tools can be installed locally on your own machine, or run via a public Web interface. Google for EMBOSS explorer, public access points include http://emboss.bioinformatics.nl/ .
Access an EMBOSS Explorer services and explore some of the tools:
Task:
- Local composition
- Find
pepinfounder the PROTEIN COMPOSITION heading. - Retrieve the YFO Mbp1 related sequence from your R database, e.g. with something like
db$protein[db$protein$name == "UMAG_1122", "sequence"] - Copy and paste the sequence into the input field.
- Run with default parameters.
- Scroll to the figures all the way at the bottom.
- Do the same in a separate window for yeast Mbp1.
- Try to compare ... (kind of hard without reference, overlay and expectation, isn't it?)
Task:
- Global composition
- Find
pepstatsunder the PROTEIN COMPOSITION heading. - Paste the YFO Mbp1 sequence in FASTA format into the input field.
- Run with default parameters.
- Do the same in a separate window for yeast Mbp1.
- Try to compare ... are there significant and unexpected differences?
Task:
- Motifs
- Find
pepcoils, an algorithm to detect coiled coil motifs. - Run this with the YFO Mbp1 sequence and yeast Mbp1.
- Try to compare ... do both sequences have coiled-coil motif predictions? Are they annotated in approximately comparable regions of the respective sequence?
Task:
- Transmembrane sequences
- Find
tmap. Also findshuffleseq. - Use your YFO sequence to annotate transmembrane helices for your protein and for a few shuffled sequences. The YFO is not expected to have TM helices, nor are the shuffled sequences expected to have any. If you do find some, these are most likely "false positives".
- Also compare the following positive control: Gef1 - a yeast chloride channel with 10 trans-membrane helices and outward localized N-terminus:
>gi|6322500|ref|NP_012574.1| Gef1p [Saccharomyces cerevisiae S288c]
MPTTYVPINQPIGDGEDVIDTNRFTNIPETQNFDQFVTIDKIAEENRPLSVDSDREFLNSKYRHYREVIW
DRAKTFITLSSTAIVIGCIAGFLQVFTETLVNWKTGHCQRNWLLNKSFCCNGVVNEVTSTSNLLLKRQEF
ECEAQGLWIAWKGHVSPFIIFMLLSVLFALISTLLVKYVAPMATGSGISEIKVWVSGFEYNKEFLGFLTL
VIKSVALPLAISSGLSVGKEGPSVHYATCCGYLLTKWLLRDTLTYSSQYEYITAASGAGVAVAFGAPIGG
VLFGLEEIASANRFNSSTLWKSYYVALVAITTLKYIDPFRNGRVILFNVTYDRDWKVQEIPIFIALGIFG
GLYGKYISKWNINFIHFRKMYLSSWPVQEVLFLATLTALISYFNEFLKLDMTESMGILFHECVKNDNTST
FSHRLCQLDENTHAFEFLKIFTSLCFATVIRALLVVVSYGARVPAGIFVPSMAVGATFGRAVSLLVERFI
SGPSVITPGAYAFLGAAATLSGITNLTLTVVVIMFELTGAFMYIIPLMIVVAITRIILSTSGISGGIADQ
MIMVNGFPYLEDEQDEEEEETLEKYTAEQLMSSKLITINETIYLSELESLLYDSASEYSVHGFPITKDED
KFEKEKRCIGYVLKRHLASKIMMQSVNSTKAQTTLVYFNKSNEELGHRENCIGFKDIMNESPISVKKAVP
VTLLFRMFKELGCKTIIVEESGILKGLVTAKDILRFKRIKYREVHGAKFTYNEALDRRCWSVIHFIIKRF
TTNRNGNVI- Evaluate the output: does the algorithm (wrongly) predict TM-helices in your protein? In the shuffled sequences? Does it find all ten TM-helices in Gef1?
Try to familiarize yourself with the offerings in the EMBOSS package. I find some of the nucleic acid tools indispensable in the lab, such as restriction-site mapping tools, and I frequently use the alignment tools Needle and Water, but by and large the utility of many of the components–while fast, efficient and straightforward to use– suffers from lack of reference and comparison and from terse output. The routines show their conceptual origin in the 1970s and 1980s. We will encounter alternatives in later assignments.
R Sequence Analysis Tools
It's interesting to see this collection of tools that were carefully designed some twenty years ago, as an open source replacement for a set of software tools - the GCG package - that was indispensable for molecular biology labs in the 80s and 90s, but whose cost had become prohibitive. Fundamentally this is a building block approach, and the field has turned to programming solutions instead.
As for functionality, much more sophisticated functions are available on the Web: do take a few minutes and browse the curated Web services directory of bioinformatics.ca.
As for versatility, R certainly has the edge. Let's explore some of the functions available in the seqinr package that you already encountered in the introductory R tutorial. They are comparatively basic - but it is easy to prepare our own analysis.
if (!require(seqinr, quietly=TRUE)) {
install.packages("seqinr")
library(seqinr)
}
help(package = seqinr) # shows the available functions
# Let's try a simple function
?computePI
# This takes as input a vector of upper-case AA codes
# Let's retrieve the YFO sequence from our datamodel
# (assuming it is the last one that was added):
db$protein[nrow(db$protein), "sequence"]
# We can use the function strsplit() to split the string
# into single characters, but we transform the sequence
# into uppercase first.
s <- db$protein[nrow(db$protein), "sequence"]
s <- toupper(s)
s <- strsplit(s, "") # splitting on the empty spring
# splits into single characters
s <- unlist(s)
# Alternatively, seqinr provides
# the function s2c() to convert strings into
# character vectors (and c2s to convert them back).
s <- s2c(toupper(db$protein[nrow(db$protein), "sequence"]))
s
computePI(s) # isoelectric point
pmw(s) # molecular weight
AAstat(s) # This also plots the distribution of
# values along the sequence
# A true Labor of Love has gone into the
# compilation of the "aaindex" data:
?aaindex
data(aaindex)
length(aaindex)
# Here are all the index descriptions
for (i in 1:length(aaindex)) {
cat(paste(i, ": ", aaindex[[i]]$D, "\n", sep=""))
}
# Lets use one of the indices to calculate and plot amino-acid
# composition enrichment:
aaindex[[459]]
<source>
=== Sequence Composition Enrichment ===
We will plot an enrichment plot to compare one of the amino acid indices with the
situation in our sequence.
<source lang="R">
refData <- aaindex[[459]]$I # reference frequencies in %
names(refData) <- a(names(refData)) # change names to single-letter
# code using seqinr's "a()" function
refData
# tabulate our sequence of interest and normalize
obsData <- table(s) # count occurrences
obsData = 100 * (obsData / sum(obsData)) # Normalize
obsData
len <- length(refData)
logRatio <- numeric() # create an empty vector
# loop over all elements of the reference, calculate log-ratios
# and store them in the vector
for (i in 1:len) {
aa <- names(refData)[i] # get the name of that amino acid
fObs <- obsData[aa] # retrieve the frequency for that name
fRef <- refData[aa]
logRatio[aa] <- log(fObs / fRef) / log(2)
}
barplot(logRatio)
# Sort by frequency, descending
logRatio <- sort(logRatio, decreasing = TRUE)
barplot(logRatio)
# label the y-axis
# (see text() for details)
label <- expression(paste(log[2],"( f(obs) / f(ref) )", sep = ""))
barplot(logRatio,
main = paste("AA composition enrichment"),
ylab = label,
cex.names=0.9)
# color the bars by type
# define colors
chargePlus <- "#282A3F"
chargeMinus <- "#531523"
hydrophilic <- "#47707F"
hydrophobic <- "#6F7A79"
plain <- "#EDF7F7"
# Assign the colors to the different amino acid names
barColors <- character(len)
for (i in 1:length(refData)) {
AA <- names(logRatio[i])
if (grepl("[HKR]", AA)) {barColors[i] <- chargePlus }
else if (grepl("[DE]", AA)) {barColors[i] <- chargeMinus}
else if (grepl("[NQST]", AA)) {barColors[i] <- hydrophilic}
else if (grepl("[FAMILYVW]", AA)) {barColors[i] <- hydrophobic}
else barColors[i] <- plain
}
barplot(logRatio,
main = paste("AA composition enrichment"),
ylab = label,
col = barColors,
cex.names=0.9)
# draw a line at 0
abline(h=0)
# add a legend that indicates what the colours mean
legend (x = 1,
y = -1,
legend = c("charged (+)",
"charged (-)",
"hydrophilic",
"hydrophobic",
"plain"),
bty = "n",
fill = c(chargePlus,
chargeMinus,
hydrophilic,
hydrophobic,
plain)
)Task:
Interpret this plot. (Can you?)
- That is all.
Links and resources
- RegEx Pal - a tool for testing and developing regular expressions.
Footnotes and references
Ask, if things don't work for you!
- If anything about the assignment is not clear to you, please ask on the mailing list. You can be certain that others will have had similar problems. Success comes from joining the conversation.
- Do consider how to ask your questions so that a meaningful answer is possible:
- How to create a Minimal, Complete, and Verifiable example on stackoverflow and ...
- How to make a great R reproducible example are required reading.
| < Assignment 2 | Assignment 4 > |