BIO Assignment Week 3
Assignment for Week 3
Sequence Analysis
| < Assignment 2 | Assignment 4 > |
Note! This assignment is currently active. All significant changes will be announced on the mailing list.
Concepts and activities (and reading, if applicable) for this assignment will be topics on next week's quiz.
Contents
Introduction
Store
The Protein datamodel
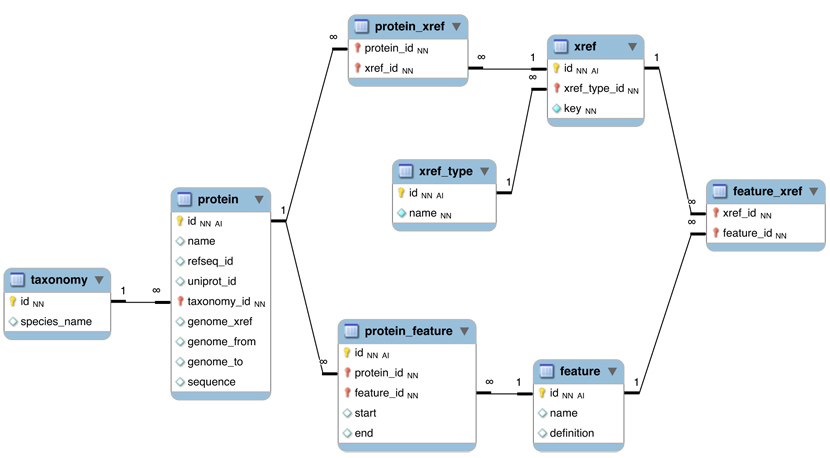
Here is a revised schema for the initial model that we developed in class:
- The
proteintable is at the centre. Its primary key is a unique integer. We store the NCBI RefSeq ID and the Uniprot ID in the table. We would not call them "foreign keys", since the information they reference is not in our schema but at the NCBI/EBI. Thegenome_xreffield will hold the ID for the actual genomic sequence, where the coding region of the gene is identified by its start ("from") and end ("to"). - The
taxonomytable holds information about species. We use the NCBI taxonomy ID as its primary key. The same key appears in the protein table as the foreign key that links the protein with its proper taxonomy information. - The
protein_featuretable links a protein with all the features that we annotate it with. Start and end coordinates identify the region of the sequence we have annotated. In principle these attributes could be NULL. - The
featuretable contains a feature name and a textual definition. Again, these attributes could theoretically be NULL. - The
protein_xreftable and thefeature_xreftable connect cross-reference data. - The
xreftable needs to hold two attributes: the type of cross-reference (e.g. PubMed), from which we would construct an URL or other accession method, and the actual key (e.g.10747782). The type of cross-reference needs some thought however: this field needs a "controlled vocabulary" to ensure that the same cross-reference is described consistently with the same string (pubmed? PubMed? Pub-Med?). There are a number of ways to achieve this, the best way is to store these types in their own table -xref_type- and link to that table via a foreign key.
Implementing the Data Model in R
To actually implement the model in R we will create the tables as data frames, and we will collect them in a list. We don't have to keep the tables in a list - we could also keep them as independent objects, but with a table named "protein" we should be worried of inadvertently overwriting the table. A list is neater, more flexible (we might want to have several of these), it reflects our intent about the model better, and doesn't require very much more typing. For now, to keep things simple, we will implement only two tables: protein and taxonomy. We'll add the rest when we actually need them.
db <- list()
db$protein <- data.frame(id = 1,
name = "Mbp1",
refseq_id = "NP_010227",
uniprot_id = "P39678",
taxonomy_id = 4932,
genome_xref = "NC_001136.10",
genome_from = 352877,
genome_to = 355378,
sequence = "...",
stringsAsFactors = FALSE)
db$taxonomy <- data.frame(id = 4932,
species_name = "Saccharomyces cerevisiae",
stringsAsFactors = FALSE)
# Let's add a second protein:
db$protein <- rbind(db$protein,
data.frame(id = 2,
name = "Res2",
refseq_id = "NP_593032",
uniprot_id = "P41412",
taxonomy_id = 4896,
genome_xref = "NC_003424.3",
genome_from = 686543,
genome_to = 689179,
sequence = "...",
stringsAsFactors = FALSE))
db$taxonomy <- rbind(db$taxonomy,
data.frame(id = 4896,
species_name = "Schizosaccharomyces pombe",
stringsAsFactors = FALSE))
str(db)
# you can look at the contents of the tables in the usual way:
db$protein
db$protein$refseq_id
db$protein[,"name"]
db$taxonomy
db$taxonomy[1,]
# What is the species name for the protein
# whose name is "Mbp1"?
# Get the taxonomy_id. This is a cell in the
# table: db$protein[<row>, <column>]
# <row> is the row in which the value in
# the name column is Mbp1:
db$protein$name == "Mbp1"
# The <column> is called "taxonomy_id". Simply
# insert these two expressions in the square
# brackets.
db$protein[db$protein$name == "Mbp1", "taxonomy_id"]
# Assign the taxonomy_id value ...
x <- db$protein[db$protein$name == "Mbp1", "taxonomy_id"]
# ... and fetch the species_name value from db$taxonomy
db$taxonomy[db$taxonomy$id == x, "species_name"]This works, but you can appreciate why we don't usually want to type all this this by hand - it's much more convenient, with less potential for error to write the necessary get- and set- functions.
We only entered a placeholder for the sequence field. Sequences come in many different flavours when we copy them from a Webpage: there can be whitespace, carriage returns, numbers, they can be upper-case, lower-case mixed-case ...
What we want in our sequence data field is one string that contains the entire sequence, and nothing but upper-case amino-acid code letters.
We'll need to look at how we work with strings in R, how we identify and work with patterns. This is a great time to introduce regular expressions.
A Brief First Encounter of Regular Expressions
- Regular expressions are a concise description language to define patterns for pattern-matching in strings.
Truth be told, many programmers have a love-hate relationship with regular expressions. The syntax of regular expressions is very powerful and expressive, but also terse, not always intuitive, and sometimes hard to understand. I'll introduce you to a few principles here that are quite straightforward and they will probably cover 99% of the cases you will encounter.
Here is our test-case: the sequence of Mbp1, copied from the Genbank Webpage.
1 msnqiysary sgvdvyefih stgsimkrkk ddwvnathil kaanfakakr trilekevlk
61 ethekvqggf gkyqgtwvpl niakqlaekf svydqlkplf dftqtdgsas pppapkhhha
121 skvdrkkair sastsaimet krnnkkaeen qfqsskilgn ptaaprkrgr pvgstrgsrr
181 klgvnlqrsq sdmgfprpai pnssisttql psirstmgpq sptlgileee rhdsrqqqpq
241 qnnsaqfkei dledglssdv epsqqlqqvf nqntgfvpqq qssliqtqqt esmatsvsss
301 pslptspgdf adsnpfeerf pgggtspiis miprypvtsr pqtsdindkv nkylsklvdy
361 fisnemksnk slpqvllhpp phsapyidap idpelhtafh wacsmgnlpi aealyeagts
421 irstnsqgqt plmrsslfhn sytrrtfpri fqllhetvfd idsqsqtvih hivkrksttp
481 savyyldvvl skikdfspqy rielllntqd kngdtalhia skngdvvffn tlvkmgaltt
541 isnkegltan eimnqqyeqm miqngtnqhv nssntdlnih vntnnietkn dvnsmvimsp
601 vspsdyityp sqiatnisrn ipnvvnsmkq masiyndlhe qhdneikslq ktlksisktk
661 iqvslktlev lkesskdeng eaqtnddfei lsrlqeqntk klrkrliryk rlikqkleyr
721 qtvllnklie detqattnnt vekdnntler lelaqeltml qlqrknklss lvkkfednak
781 ihkyrriire gtemnieevd ssldvilqtl iannnknkga eqiitisnan sha
//
Task:
Navigate to http://regexpal.com and paste the sequence into the lower box. This site is one of a few online regular expression testers and invaluable when you are developing regular expression patterns.
Lets try some expressions:
- Most characters are matched literally.
- Type "
a" in to the upper box and you will see all "a" characters matched. Then replaceawithq. - Now type "
aa" instead. Thenkrnnkk. Sequences of charcters are also matched literally.
- We can specify more than one character to match if we place it in square brackets.
[lq]matcheslORq.[familyvw]matches hydrophobic amino acids.
- Within square brackets, we can specify "ranges".
[1-5]matches digits from 1 to 5.
- Within square brackets, we can specify characters that should NOT be matched, with the caret,
^. [^0-9]matches everything EXCEPT digits.[^a-z]matches everything that is not a lower-case letter.
One of the R functions that uses regular expressions is the function gsub() that replaces characters that match a "regex" with other characters. That is useful for our purpose: we can
- match to all characters that are NOT a letter, and
- replace them by - nothing: the empty string
"".
This deletes them.
Task:
Try this code:
seq <- "
1 msnqiysary sgvdvyefih stgsimkrkk ddwvnathil kaanfakakr trilekevlk
61 ethekvqggf gkyqgtwvpl niakqlaekf svydqlkplf dftqtdgsas pppapkhhha
121 skvdrkkair sastsaimet krnnkkaeen qfqsskilgn ptaaprkrgr pvgstrgsrr
181 klgvnlqrsq sdmgfprpai pnssisttql psirstmgpq sptlgileee rhdsrqqqpq
241 qnnsaqfkei dledglssdv epsqqlqqvf nqntgfvpqq qssliqtqqt esmatsvsss
301 pslptspgdf adsnpfeerf pgggtspiis miprypvtsr pqtsdindkv nkylsklvdy
361 fisnemksnk slpqvllhpp phsapyidap idpelhtafh wacsmgnlpi aealyeagts
421 irstnsqgqt plmrsslfhn sytrrtfpri fqllhetvfd idsqsqtvih hivkrksttp
481 savyyldvvl skikdfspqy rielllntqd kngdtalhia skngdvvffn tlvkmgaltt
541 isnkegltan eimnqqyeqm miqngtnqhv nssntdlnih vntnnietkn dvnsmvimsp
601 vspsdyityp sqiatnisrn ipnvvnsmkq masiyndlhe qhdneikslq ktlksisktk
661 iqvslktlev lkesskdeng eaqtnddfei lsrlqeqntk klrkrliryk rlikqkleyr
721 qtvllnklie detqattnnt vekdnntler lelaqeltml qlqrknklss lvkkfednak
781 ihkyrriire gtemnieevd ssldvilqtl iannnknkga eqiitisnan sha
//
"
seq # "\n" means: line-break
seq <- gsub("[^a-zA-Z]", "", seq)
seq
# Now to change the sequence to upper-case. R has toupper()
# and tolower().
toupper(seq)
# Lets put this in a function:
sanitizeSequence <- function(s) {
return(toupper(gsub("[^a-zA-Z]", "", s)))
}
# try it:
test <- "f123 a$^&&*)m. i l马马虎虎 é yßv+w"
sanitizeSequence(test)A Function to Set Values
Building on this function, we can write a function to set values in our protein table.
setDB <- function(database,
table,
id,
name = "",
refseq_id = "",
uniprot_id = "",
taxonomy_id = "",
genome_xref = "",
genome_from = "",
genome_to = "",
sequence = "",
species_name = "") {
if (missing(database) | missing(table) | missing(id)) {
stop("database, table and id have to be specified")
}
if (table == "protein") {
row <- which(database$protein$id == id)
if (name != "") { database$protein[row, "name"] <- name }
if (refseq_id != "") { database$protein[row, "refseq_id"] <- refseq_id}
if (uniprot_id != "") { database$protein[row, "uniprot_id"] <- uniprot_id}
if (taxonomy_id != "") { database$protein[row, "taxonomy_id"] <- taxonomy_id}
if (genome_xref != "") { database$protein[row, "genome_xref"] <- genome_xref}
if (genome_from != "") { database$protein[row, "genome_from"] <- genome_from}
if (genome_to != "") { database$protein[row, "genome_to"] <- genome_to}
if (sequence != "") { database$protein[row, "sequence"] <- sanitizeSequence(sequence)}
}
if (table == "taxonomy") {
row <- which(database$taxonomy$id == id)
if (species_name != "") { database$taxonomy[row, "species_name"] <- species_name }
}
return(database)
}
# Note that this code could be significantly improved: we are
# not doing any tests that the values are sane.
# But for now I just want to keep the code simple.
db$protein[1,]
db <- setDB(database = db, table="protein", id=1, name="Armadillo")
db$protein[1,]
db <- setDB(database = db, table="protein", id=1, name="Mbp1")
# With this, we can update our entries to set the sequence to its actual value.
seq
seq <- "
1 msnqiysary sgvdvyefih stgsimkrkk ddwvnathil kaanfakakr trilekevlk
61 ethekvqggf gkyqgtwvpl niakqlaekf svydqlkplf dftqtdgsas pppapkhhha
121 skvdrkkair sastsaimet krnnkkaeen qfqsskilgn ptaaprkrgr pvgstrgsrr
181 klgvnlqrsq sdmgfprpai pnssisttql psirstmgpq sptlgileee rhdsrqqqpq
241 qnnsaqfkei dledglssdv epsqqlqqvf nqntgfvpqq qssliqtqqt esmatsvsss
301 pslptspgdf adsnpfeerf pgggtspiis miprypvtsr pqtsdindkv nkylsklvdy
361 fisnemksnk slpqvllhpp phsapyidap idpelhtafh wacsmgnlpi aealyeagts
421 irstnsqgqt plmrsslfhn sytrrtfpri fqllhetvfd idsqsqtvih hivkrksttp
481 savyyldvvl skikdfspqy rielllntqd kngdtalhia skngdvvffn tlvkmgaltt
541 isnkegltan eimnqqyeqm miqngtnqhv nssntdlnih vntnnietkn dvnsmvimsp
601 vspsdyityp sqiatnisrn ipnvvnsmkq masiyndlhe qhdneikslq ktlksisktk
661 iqvslktlev lkesskdeng eaqtnddfei lsrlqeqntk klrkrliryk rlikqkleyr
721 qtvllnklie detqattnnt vekdnntler lelaqeltml qlqrknklss lvkkfednak
781 ihkyrriire gtemnieevd ssldvilqtl iannnknkga eqiitisnan sha
//
"
db <- setDB(database = db, table="protein", id=1, sequence=seq)
seq <- "
1 maprssavhv avysgvevye cfikgvsvmr rrrdswlnat qilkvadfdk pqrtrvlerq
61 vqigahekvq ggygkyqgtw vpfqrgvdla tkykvdgims pilsldideg kaiapkkkqt
121 kqkkpsvrgr rgrkpsslss stlhsvnekq pnssisptie ssmnkvnlpg aeeqvsatpl
181 paspnallsp ndntikpvee lgmleapldk yeeslldffl hpeegripsf lyspppdfqv
241 nsvidddght slhwacsmgh iemiklllra nadigvcnrl sqtplmrsvi ftnnydcqtf
301 gqvlellqst iyavdtngqs ifhhivqsts tpskvaaaky yldcilekli siqpfenvvr
361 lvnlqdsngd tslliaarng amdcvnslls ynanpsipnr qrrtaseyll eadkkphsll
421 qsnsnashsa fsfsgispai ispscsshaf vkaipsissk fsqlaeeyes qlrekeedli
481 ranrlkqdtl neisrtyqel tflqknnpty sqsmenlire aqetyqqlsk rlliwlearq
541 ifdlerslkp htslsisfps dflkkedgls lnndfkkpac nnvtnsdeye qlinkltslq
601 asrkkdtlyi rklyeelgid dtvnsyrrli amscginped lsleildave ealtrek
"
db <- setDB(database = db, table="protein", id=2, sequence=seq)
A Function to Add New Entries
The function to add a new entry to the database looks quite similar. But we have to add a test whether the species name has been entered if it's a new taxonomy ID.
addToDB <- function(database,
name = "",
refseq_id = "",
uniprot_id = "",
taxonomy_id = "",
genome_xref = "",
genome_from = "",
genome_to = "",
sequence = "",
species_name = "") {
if (missing(database) | missing(taxonomy_id)) {
stop("database and taxonomy_id have to be specified")
}
# handle taxonomy
if (! any(db$taxonomy$id == taxonomy_id)) { # new taxonomy_id
if (missing(species_name)) {
stop("taxonomy_id is new: species_name has to be specified")
}
else {
# add this species to the taxonomy table
database$taxonomy <- rbind(database$taxonomy,
data.frame(id = taxonomy_id,
species_name = species_name,
stringsAsFactors = FALSE))
}
}
# handle protein
# get a new protein id
pid <- max(database$protein$id) + 1
database$protein <- rbind(db$protein,
data.frame(id = pid,
name = name,
refseq_id = refseq_id,
uniprot_id = uniprot_id,
taxonomy_id = taxonomy_id,
genome_xref = genome_xref,
genome_from = genome_from,
genome_to = genome_to,
sequence = sanitizeSequence(sequence),
stringsAsFactors = FALSE))
return(database)
}
# Test this.
newSeq <- "
1 msgdktifka tysgvpvyec iinnvavmrr rsddwlnatq ilkvvgldkp qrtrvlerei
61 qkgihekvqg gygkyqgtwi pldvaielae ryniqgllqp itsyvpsaad spppapkhti
121 stsnrskkii padpgalgrs rratsietes evigaapnnv segsmspsps dissssrtps
181 plpadrahpl hanhalagyn grdannhary adiildyfvt enttvpslli npppdfnpdm
241 sidddehtal hwacamgrir vvklllsaga difrvnsnqq talmratmfs nnydlrkfpe
301 lfellhrsil nidrndrtvf hhvvdlalsr gkphaaryym etminrlady gdqladilnf
361 qddegetplt maararskrl vrlllehgad pkirnkegkn aedyiieder frsspsrtgp
421 agielgadgl pvlptsslht seagqrtagr avtlmsnllh sladsydsei ntaekkltqa
481 hgllkqiqte iedsakvaea lhheaqgvde erkrvdslql alkhainkra rddlerrwse
541 gkqaikrarl qaglepgals tsnatnapat gdqkskddak sliealpagt nvktaiaelr
601 kqlsqvqank telvdkfvar areqgtgrtm aayrrliaag cggiapdevd avvgvlcell
661 qeshtgarag aggerddrar dvammlkgag aaalaanaga p
"
db2 <- addToDB(db,
name = "UMAG_1122",
refseq_id = "XP_011392621",
uniprot_id = "A0A0D1DP35",
taxonomy_id = "5270",
genome_xref = "NC_026499.1",
genome_from = "150555",
genome_to = "152739",
sequence = newSeq,
species_name = "Ustilago maydis")As you can see, adding a new protein to our database is now rather compact.
Task:
In the last assignment, you discovered a protein that is related to yaest Mbp1 in YFO.
Add it to the database. Ask on the mailing list if you don't know how (but be specific) or if you don't know how to find particular information items.
- TBC
Links and resources
Footnotes and references
Ask, if things don't work for you!
- If anything about the assignment is not clear to you, please ask on the mailing list. You can be certain that others will have had similar problems. Success comes from joining the conversation.
- Do consider how to ask your questions so that a meaningful answer is possible:
- How to create a Minimal, Complete, and Verifiable example on stackoverflow and ...
- How to make a great R reproducible example are required reading.
| < Assignment 2 | Assignment 4 > |