BIO Assignment Week 8
Assignment for Week 7
Multiple Sequence Alignment
Note! This assignment is currently inactive. Major and minor unannounced changes may be made at any time.
Concepts and activities (and reading, if applicable) for this assignment will be topics on next week's quiz.
Contents
Introduction
In the last assignment we discovered homologs to S. cerevisiae Mbp1 in YFO. Some of these will be orthologs to Mbp1, some will be paralogs. Some will have similar function, some will not. We discussed previously that genes that evolve under continuously similar evolutionary pressure should be most similar in sequence, and should have the most similar "function".
In this assignment we will define the YFO gene that is the most similar ortholog to S. cerevisiae Mbp1, and perform a multiple sequence alignment with it.
Let us briefly review the basic concepts.
Orthologs and Paralogs revisited
- All related genes are homologs.
Two central definitions about the mutual relationships between related genes go back to Walter Fitch who stated them in the 1970s:
- Orthologs have diverged after speciation.
- Paralogs have diverged after duplication.
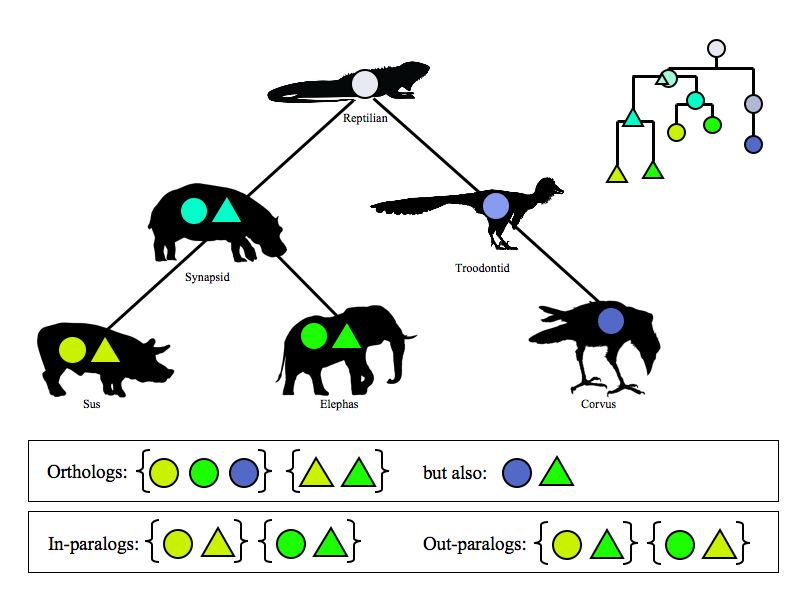
Defining orthologs
To be reasonably certain about orthology relationships, we would need to construct and analyze detailed evolutionary trees. This is computationally expensive and the results are not always unambiguous either, as we will see in a later assignment. But a number of different strategies are available that use precomputed results to define orthologs. These are especially useful for large, cross genome surveys. They are less useful for detailed analysis of individual genes. Pay the sites a visit and try a search.
- Orthologs by COGs and KOGS
- The COGs database is a database of clusters of mutually orthologous bacterial genes at the NCBI and the KOGS database is its eukaryotic counterpart. I am sure it is very well done, but the interface is useless. One could download the data and code one's own analysis routines I guess...
- Orthologs by OMA and OrthoDB
- OrthoDB includes a large number of species, among them all of our protein-sequenced fungi. However the search function (by keyword) retrieves many paralogs together with the orthologs, for example, the yeast Swi4 protein is found together with yeast Mbp1 and these two are clearly paralogs.
- OMA (the Orthologous Matrix) maintained at the Swiss Federal Institute of Technology contains a large number of orthologs from sequenced genomes. Searching with
MBP1_YEAST(this is the Swissprot ID) as a "Group" search finds the correct gene in EREGO, KLULA, CANGL and SACCE. But searching with the sequence of the Ustilago maydis ortholog does not find the yeast protein, but the orthologs in YARLI, SCHPO, LACCBI, CRYNE and USTMA. Apparently the orthologous group has been split into several subgroups across the fungi.
- Orthologs by syntenic gene order conservation
- We will revisit this when we explore the UCSC genome browser.
- Orthologs by RBM
- This is easy to do. Simply pick the gene which you have identified and annotated for YFO in Assignment 3 and confirm that it is the best match in yeast. The results are unambiguous regarding the applied criterion, but there may be residual doubt whether these two best-matching sequences are actually the most similar orthologs.
Task:
- Navigate to the BLAST homepage.
- Paste the YFO RefSeq sequence identifier into the search field. (You don't have to search with sequences–you can search directly with an NCBI identifier IF you want to search with the full-length sequence.)
- Set the database to refseq, and restrict the species to Saccharomyces cerevisiae.
- Run BLAST.
- Keep the window open for the next task.
The top hit should be yeast Mbp1 (NP_010227). E mail me your sequence identifiers if it is not. If it is, you have confirmed the RBM or BBM criterion (Reciprocal Best Match or Bidirectional Best Hit, respectively).
Technically, this is not perfectly true since you have searched with the APSES domain in one direction, with the full-length sequence in the other. For this task I wanted you to try the search-with-accession-number. Therefore the procedural laxness, I hope it is permissible. In fact, performing the reverse search with the YFO APSES domain should actually be more stringent, i.e. if you find the right gene with the longer sequence, you are even more likely to find the right gene with the shorter one.
- Orthology by annotation
- The NCBI precomputes BLAST results and makes them available at the RefSeq database entry for your protein.
Task:
- In your BLAST result page, click on the RefSeq link for your query to navigate to the RefSeq database entry for your protein.
- Follow the Blink link in the right-hand column under Related information.
- Restrict the view to Fungi and RefSeq under the "Display options"
You should see a number of genes with low E-values and high coverage in other fungi - however this search is problematic since the full length gene across the database finds mostly Ankyrin domains.
You will find that all of these approaches yield some of the orthologs. But none finds them all. The take home message is: precomputed results are good for large-scale survey-type investigations, where you can't humanly process the information by hand. But for more detailed questions, careful manual searches are still indsipensable.
- Orthology by crowdsourcing
- Luckily a crowd of willing hands has prepared the necessary sequences for you: in the section below you will find a link to the annotated and verified Mbp1 orthologs from last year's course :-)
We could call this annotation by many hands "crowdsourcing" - handing out small parcels of work to many workers, who would typically allocate only a small share of their time, but here the strength is in numbers and especially projects that organize via the Internet can tally up very impressive manpower, for free, or as Microwork. These developments have some interest for bioinformatics: many of our more difficult tasks can not be easily built into an algorithm, language related tasks such as text-mining, or pattern matching tasks come to mind. Allocating this to a large number of human contributors may be a viable alternative to computation. A marketplace where this kind of work is already a reality is Amazon's "Mechanical Turk" Marketplace: programmers–"requesters"– use an open interface to post tasks for payment, "providers" from all over the world can engage in these. Tasks may include matching of pictures, or evaluating the aesthetics of competing designs. A quirky example I came across recently was when information designer David McCandless had 200 "Mechanical Turks" draw a small picture of their soul for his collection.
The name "Mechanical Turk" by the way relates to a famous ruse, when a Hungarian inventor and adventurer toured the imperial courts of 18th century Europe with an automaton, dressed in turkish robes and turban, that played chess at the grandmaster level against opponents that included Napoleon Bonaparte and Benjamin Franklin. No small mechanical feat in any case, it was only in the 19th century that it was revealed that the computational power was actually provided by a concealed human.
Are you up for some "Turking"? Before the next quiz, edit the Mbp1 RBM page on the Student Wiki and include the RBM for Mbp1, for a 10% bonus on the next quiz.
Align and Annotate
Review of domain annotations
APSES domains are relatively easy to identify and annotate but we have had problems with the ankyrin domains in Mbp1 homologues. Both CDD as well as SMART have identified such domains, but while the domain model was based on the same Pfam profile for both, and both annotated approximately the same regions, the details of the alignments and the extent of the predicted region was different.
Mbp1 forms heterodimeric complexes with a homologue, Swi6. Swi6 does not have an APSES domain, thus it does not bind DNA. But it is similar to Mbp1 in the region spanning the ankyrin domains and in 1999 Foord et al. published its crystal structure (1SW6). This structure is a good model for Ankyrin repeats in Mbp1. For details, please refer to the consolidated Mbp1 annotation page I have prepared.
In what follows, we will use the program JALVIEW - a Java based multiple sequence alignment editor to load and align sequences and to consider structural similarity between yeast Mbp1 and its closest homologue in your organism.
In this part of the assignment,
- You will load sequences that are most similar to Mbp1 into an MSA editor;
- You will add sequences of ankyrin domain models;
- You will perform a multiple sequence alignment;
- You will try to improve the alignment manually;
Jalview, loading sequences
Geoff Barton's lab in Dundee has developed an integrated MSA editor and sequence annotation workbench with a number of very useful functions. It is written in Java and should run on Mac, Linux and Windows platforms without modifications.
| Waterhouse et al. (2009) Jalview Version 2--a multiple sequence alignment editor and analysis workbench. Bioinformatics 25:1189-91. (pmid: 19151095) |
|
[ PubMed ] [ DOI ] UNLABELLED: Jalview Version 2 is a system for interactive WYSIWYG editing, analysis and annotation of multiple sequence alignments. Core features include keyboard and mouse-based editing, multiple views and alignment overviews, and linked structure display with Jmol. Jalview 2 is available in two forms: a lightweight Java applet for use in web applications, and a powerful desktop application that employs web services for sequence alignment, secondary structure prediction and the retrieval of alignments, sequences, annotation and structures from public databases and any DAS 1.53 compliant sequence or annotation server. AVAILABILITY: The Jalview 2 Desktop application and JalviewLite applet are made freely available under the GPL, and can be downloaded from www.jalview.org. |
We will use this tool for this assignment and explore its features as we go along.
Task:
- Navigate to the Jalview homepage click on Download, install Jalview on your computer and start it. A number of windows that showcase the program's abilities will load, you can close these.
- Prepare homologous Mbp1 sequences for alignment:
- Open the All Mbp1 proteins page. (This is the list of Mbp1 orthologs I mentioned above.)
- Copy the FASTA sequences of the reference proteins, paste them into a text file (TextEdit on the Mac, Notepad on Windows) and save the file; you could give it an extension of
.fa–but you don't have to. - Check whether the sequence for YFO is included in the list. If it is, fine. If it is not, retrieve it from NCBI, paste it into the file and edit the header like the other sequences. If the wrong sequence from YFO is included, replace it and let me know.
- Return to Jalview and select File → Input Alignment → from File and open your file. A window with sequences should appear.
- Copy the sequences for ankyrin domain models (below), click on the Jalview window, select File → Add sequences → from Textbox and paste them into the Jalview textbox. Paste two separate copies of the CD00204 consensus sequence and one copy of 1SW6.
- When all the sequences are present, click on Add.
Jalview now displays all the sequences, but of course this is not yet an alignment.
- Ankyrin domain models
>CD00204 ankyrin repeat consensus sequence from CDD NARDEDGRTPLHLAASNGHLEVVKLLLENGADVNAKDNDGRTPLHLAAKNGHLEIVKLLL EKGADVNARDKDGNTPLHLAARNGNLDVVKLLLKHGADVNARDKDGRTPLHLAAKNGHL
>1SW6 from PDB - unstructured loops replaced with xxxx GPIITFTHDLTSDFLSSPLKIMKALPSPVVNDNEQKMKLEAFLQRLLFxxxxSFDSLLQE VNDAFPNTQLNLNIPVDEHGNTPLHWLTSIANLELVKHLVKHGSNRLYGDNMGESCLVKA VKSVNNYDSGTFEALLDYLYPCLILEDSMNRTILHHIIITSGMTGCSAAAKYYLDILMGW IVKKQNRPIQSGxxxxDSILENLDLKWIIANMLNAQDSNGDTCLNIAARLGNISIVDALL DYGADPFIANKSGLRPVDFGAG
Computing alignments
The EBI has a very convenient page to access a number of MSA algorithms. This is especially convenient when you want to compare, e.g. T-Coffee and Muscle and MAFFT results to see which regions of your alignment are robust. You could use any of these tools, just paste your sequences into a Webform, download the results and load into Jalview. Easy.
But even easier is to calculate the alignments directly from Jalview. Or at least it is, when the service is actually available. (Not today. Bummer.)
- Calculate a MAFFT alignment when the Jalview Web service is available
Task:
- In Jalview, select Web Service → Alignment → MAFFT Multiple Protein Sequence Alignment. The alignment is calculated in a few minutes and displayed in a new window.
- Calculate a MAFFT alignment when the Jalview Web service is NOT available
Task:
- In Jalview, select File → Output to Textbox → FASTA
- Copy the sequences.
- Navigate to the MAFFT Input form at the EBI.
- Paste your sequences into the form.
- Click on Submit.
- Close the Jalview sequence window and either save your MAFFT alignment to file and load in Jalview, or simply 'File → Input Alignment → from Textbox, paste and click New Window.
In any case, you should now have an alignment.
Task:
- Choose Colour → Hydrophobicity and → by Conservation. Then select Modify Conservation Threshold... and adjust the slider left or right to see which columns are highly conserved. You will notice that the Swi6 sequence that was supposed to align only to the ankyrin domains was in fact aligned to other parts of the sequence as well. This is one part of the MSA that we will have to correct manually and a common problem when aligning sequences of different lengths.
Editing ankyrin domain alignments
A good MSA comprises only columns of residues that play similar roles in the proteins' mechanism and/or that evolve in a comparable structural context. Since the alignment reflects the result of biological selection and conservation, it has relatively few indels and the indels it has are usually not placed into elements of secondary structure or into functional motifs. The contiguous features annotated for Mbp1 are expected to be left intact by a good alignment.
A poor MSA has many errors in its columns; these contain residues that actually have different functions or structural roles, even though they may look similar according to a (pairwise!) scoring matrix. A poor MSA also may have introduced indels in biologically irrelevant positions, to maximize spurious sequence similarities. Some of the features annotated for Mbp1 will be disrupted in a poor alignment and residues that are conserved may be placed into different columns.
Often errors or inconsistencies are easy to spot, and manually editing an MSA is not generally frowned upon, even though this is not a strictly objective procedure. The main goal of manual editing is to make an alignment biologically more plausible. Most comonly this means to mimize the number of rare evolutionary events that the alignment suggests and/or to emphasize conservation of known functional motifs. Here are some examples for what one might aim for in manually editing an alignment:
- Reduce number of indels
From a Probcons alignment: 0447_DEBHA ILKTE-K-T---K--SVVK ILKTE----KTK---SVVK 9978_GIBZE MLGLN-PGLKEIT--HSIT MLGLNPGLKEIT---HSIT 1513_CANAL ILKTE-K-I---K--NVVK ILKTE----KIK---NVVK 6132_SCHPO ELDDI-I-ESGDY--ENVD ELDDI-IESGDY---ENVD 1244_ASPFU ----N-PGLREIC--HSIT -> ----NPGLREIC---HSIT 0925_USTMA LVKTC-PALDPHI--TKLK LVKTCPALDPHI---TKLK 2599_ASPTE VLDAN-PGLREIS--HSIT VLDANPGLREIS---HSIT 9773_DEBHA LLESTPKQYHQHI--KRIR LLESTPKQYHQHI--KRIR 0918_CANAL LLESTPKEYQQYI--KRIR LLESTPKEYQQYI--KRIR
Gaps marked in red were moved. The sequence similarity in the alignment does not change considerably, however the total number of indels in this excerpt is reduced to 13 from the original 22
- Move indels to more plausible position
From a CLUSTAL alignment: 4966_CANGL MKHEKVQ------GGYGRFQ---GTW MKHEKVQ------GGYGRFQ---GTW 1513_CANAL KIKNVVK------VGSMNLK---GVW KIKNVVK------VGSMNLK---GVW 6132_SCHPO VDSKHP-----------QID---GVW -> VDSKHPQ-----------ID---GVW 1244_ASPFU EICHSIT------GGALAAQ---GYW EICHSIT------GGALAAQ---GYW
The two characters marked in red were swapped. This does not change the number of indels but places the "Q" into a a column in which it is more highly conserved (green). Progressive alignments are especially prone to this type of error.
- Conserve motifs
From a CLUSTAL alignment: 6166_SCHPO --DKRVA---GLWVPP --DKRVA--G-LWVPP XBP1_SACCE GGYIKIQ---GTWLPM GGYIKIQ--G-TWLPM 6355_ASPTE --DEIAG---NVWISP -> ---DEIA--GNVWISP 5262_KLULA GGYIKIQ---GTWLPY GGYIKIQ--G-TWLPY
The first of the two residues marked in red is a conserved, solvent exposed hydrophobic residue that may mediate domain interactions. The second residue is the conserved glycine in a beta turn that cannot be mutated without structural disruption. Changing the position of a gap and insertion in one sequence improves the conservation of both motifs.
The Ankyrin domains are quite highly diverged, the boundaries not well defined and not even CDD, SMART and SAS agree on the precise annotations. We expect there to be alignment errors in this region. Nevertheless we would hope that a good alignment would recognize homology in that region and that ideally the required indels would be placed between the secondary structure elements, not in their middle. But judging from the sequence alignment alone, we cannot judge where the secondary structure elements ought to be. You should therefore add the following "sequence" to the alignment; it contains exactly as many characters as the Swi6 sequence above and annotates the secondary structure elements. I have derived it from the 1SW6 structure
>SecStruc 1SW6 E: strand t: turn H: helix _: irregular _EEE__tt___ttt______EE_____t___HHHHHHHHHHHHHHHH_xxxx_HHHHHHH HHHH_t_____t_____t____HHHHHHH__tHHHHHHHHH____t___tt____HHHHH HH__HHHH___HHHHHHHHHHHHHEE_t____HHHHHHHHH__t__HHHHHHHHHHHHHH HHHHHH__EEE_xxxx_HHHHHt_HHHHHHH______t____HHHHHHHH__HHHHHHHH H____t____t____HHHH___
To proceed:
- Manually align the Swi6 sequence with yeast Mbp1
- Bring the Secondary structure annotation into its correct alignment with Swi6
- Bring both CDD ankyrin profiles into the correct alignment with yeast Mbp1
Proceed along the following steps:
Task:
- Add the secondary structure annotation to the sequence alignment in Jalview. Copy the annotation, select File → Add sequences → from Textbox and paste the sequence.
- Select Help → Documentation and read about Editing Alignments, Cursor Mode and Key strokes.
- Click on the yeast Mbp1 sequence row to select the entire row. Then use the cursor key to move that sequence down, so it is directly above the 1SW6 sequence. Select the row of 1SW6 and use shift/mouse to move the sequence elements and edit the alignment to match yeast Mbp1. Refer to the alignment given in the Mbp1 annotation page for the correct alignment.
- Align the secondary structure elements with the 1SW6 sequence: Every character of 1SW6 should be matched with either E, t, H, or _. The result should be similar to the Mbp1 annotation page. If you need to insert gaps into all sequences in the alignment, simply drag your mouse over all row headers - movement of sequences is constrained to selected regions, the rest is locked into place to prevent inadvertent misalignments. Remember to save your project from time to time: File → save so you can reload a previous state if anything goes wrong and can't be fixed with Edit → Undo.
- Finally align the two CD00204 consensus sequences to their correct positions (again, refer to the Mbp1 annotation page).
- You can now consider the principles stated above and see if you can improve the alignment, for example by moving indels out of regions of secondary structure if that is possible without changing the character of the aligned columns significantly. Select blocks within which to work to leave the remaining alignment unchanged. So that this does not become tedious, you can restrict your editing to one Ankyrin repeat that is structurally defined in Swi6. You may want to open the 1SW6 structure in VMD to define the boundaries of one such repeat. You can copy and paste sections from Jalview into your assignment for documentation or export sections of the alignment to HTML (see the example below).
Editing ankyrin domain alignments - Sample
This sample was created by
- Editing the alignments as described above;
- Copying a block of aligned sequence;
- Pasting it To New Alignment;
- Colouring the residues by Hydrophobicity and setting the colour saturation according to Conservation;
- Choosing File → Export Image → HTML and pasting the resulting HTML source into this Wikipage.
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
- Aligned sequences before editing. The algorithm has placed gaps into the Swi6 helix
LKWIIANand the four-residue gaps before the block of well aligned sequence on the right are poorly supported.
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
- Aligned sequence after editing. A significant cleanup of the frayed region is possible. Now there is only one insertion event, and it is placed into the loop that connects two helices of the 1SW6 structure.
Final analysis
Task:
- Compare the distribution of indels in the ankyrin repeat regions of your alignments.
- Review whether the indels in this region are concentrated in segments that connect the helices, or if they are more or less evenly distributed along the entire region of similarity.
- Think about whether the assertion that indels should not be placed in elements of secondary structure has merit in your alignment.
- Recognize that an indel in an element of secondary structure could be interpreted in a number of different ways:
- The alignment is correct, the annotation is correct too: the indel is tolerated in that particular case, for example by extending the length of an α-helix or β-strand;
- The alignment algorithm has made an error, the structural annotation is correct: the indel should be moved a few residues;
- The alignment is correct, the structural annotation is wrong, this is not a secondary structure element after all;
- Both the algorithm and the annotation are probably wrong, but we have no data to improve the situation.
(NB: remember that the structural annotations have been made for the yeast protein and might have turned out differently for the other proteins...)
You should be able to analyse discrepancies between annotation and expectation in a structured and systematic way. In particular if you notice indels that have been placed into an annotated region of secondary structure, you should be able to comment on whether the location of the indel has strong support from aligned sequence motifs, or whether the indel could possibly be moved into a different location without much loss in alignment quality.
- Considering the whole alignment and your experience with editing, you should be able to state whether the position of indels relative to structural features of the ankyrin domains in your organism's Mbp1 protein is reliable. That would be the result of this task, in which you combine multiple sequence and structural information.
- You can also critically evaluate database information that you have encountered:
- Navigate to the CDD annotation for yeast Mbp1.
- You can check the precise alignment boundaries of the ankyrin domains by clicking on the (+) icon to the left of the matching domain definition.
- Confirm that CDD extends the ankyrin domain annotation beyond the 1SW6 domain boundaries. Given your assessment of conservation in the region beyond the structural annotation: do you think that extending the annotation is reasonable also in YFO's protein? Is there evidence for this in the alignment of the CD00204 consensus with well aligned blocks of sequence beyond the positions that match Swi6?
R code: load alignment and compute information scores
As discussed in the lecture, Shannon information is calculated as the difference between expected and observed entropy, where entropy is the negative sum over probabilities times the log of those probabilities:
Here we compute Shannon information scores for aligned positions of the APSES domain, and plot the values in R. You can try this with any part of your alignment, but I have used only the aligned residues for the APSES domain for my example. This is a good choice for a first try, since there are (almost) no gaps.
Task:
- Export only the sequences of the aligned APSES domains to a file on your computer, in FASTA format. You could call this:
Mbp1_All_APSES.fa. - Explore the R-code below. Be sure that you understand it correctly. Note that there is no sampling bias correction, so positions with large numbers of gaps will receive artificially high scores.
# CalculateInformation.R
# Calculate Shannon information for positions in a multiple sequence alignment.
# Requires: an MSA in multi FASTA format
# It is good practice to set variables you might want to change
# in a header block so you don't need to hunt all over the code
# for strings you need to update.
#
setwd("/your/R/working/directory")
mfa <- "MBP1_All_APSES.fa"
# ================================================
# Read sequence alignment fasta file
# ================================================
# read MFA datafile using seqinr function read.fasta()
library(seqinr)
tmp <- read.alignment(mfa, format="fasta")
MSA <- as.matrix(tmp) # convert the list into a characterwise matrix
# with appropriate row and column names using
# the seqinr function as.matrix.alignment()
# You could have a look under the hood of this
# function to understand beter how to convert a
# list into something else ... simply type
# "as.matrix.alignment" - without the parentheses
# to retrieve the function source code (as for any
# function btw).
### Explore contents of and access to the matrix of sequences
MSA
MSA[1,]
MSA[,1]
MSA["MBP1_SACCE/1-75",1:10]
length(MSA[,1])
# ================================================
# define function to calculate entropy
# ================================================
entropy <- function(v) { # calculate shannon entropy for the aa vector v
# Note: we are not correcting for small sample sizes
# here. Thus if there are a large number of gaps in
# the alignment, this will look like small entropy
# since only a few amino acids are present. In the
# extreme case: if a position is only present in
# one sequence, that one amino acid will be treated
# as 100% conserved - zero entropy. Sampling error
# corrections are discussed eg. in Schneider et al.
# (1986) JMB 188:414
l <- length(v)
a <- rep(0, 21) # initialize a vector with 21 elements (20 aa plus gap)
# the set the name of each row to the one letter
# code. Through this, we can access a row by its
# one letter code.
names(a) <- unlist(strsplit("acdefghiklmnpqrstvwy-", ""))
for (i in 1:l) { # for the whole vector of amino acids
c <- v[i] # retrieve the character
a[c] <- a[c] + 1 # increment its count by one
} # note: we could also have used the table() function for this
tot <- sum(a) - a["-"] # calculate number of observed amino acids
# i.e. subtract gaps
a <- a/tot # frequency is observations of one amino acid
# divided by all observations. We assume that
# frequency equals probability.
a["-"] <- 0
for (i in 1:length(a)) {
if (a[i] != 0) { # if a[i] is not zero, otherwise leave as is.
# By definition, 0*log(0) = 0 but R calculates
# this in parts and returns NaN for log(0).
a[i] <- a[i] * (log(a[i])/log(2)) # replace a[i] with
# p(i) log_2(p(i))
}
}
return(-sum(a)) # return Shannon entropy
}
# ================================================
# calculate entropy for reference distribution
# (from UniProt, c.f. Assignment 2)
# ================================================
refData <- c(
"A"=8.26,
"Q"=3.93,
"L"=9.66,
"S"=6.56,
"R"=5.53,
"E"=6.75,
"K"=5.84,
"T"=5.34,
"N"=4.06,
"G"=7.08,
"M"=2.42,
"W"=1.08,
"D"=5.45,
"H"=2.27,
"F"=3.86,
"Y"=2.92,
"C"=1.37,
"I"=5.96,
"P"=4.70,
"V"=6.87
)
### Calculate the entropy of this distribution
H.ref <- 0
for (i in 1:length(refData)) {
p <- refData[i]/sum(refData) # convert % to probabilities
H.ref <- H.ref - (p * (log(p)/log(2)))
}
# ================================================
# calculate information for each position of
# multiple sequence alignment
# ================================================
lAli <- dim(MSA)[2] # length of row in matrix is second element of dim(<matrix>).
I <- rep(0, lAli) # initialize result vector
for (i in 1:lAli) {
I[i] = H.ref - entropy(MSA[,i]) # I = H_ref - H_obs
}
### evaluate I
I
quantile(I)
hist(I)
plot(I)
# you can see that we have quite a large number of columns with the same,
# high value ... what are these?
which(I > 4)
MSA[,which(I > 4)]
# And what is in the columns with low values?
MSA[,which(I < 1.5)]
# ===================================================
# plot the information
# (c.f. Assignment 5, see there for explanations)
# ===================================================
IP <- (I-min(I))/(max(I) - min(I) + 0.0001)
nCol <- 15
IP <- floor(IP * nCol) + 1
spect <- colorRampPalette(c("#DD0033", "#00BB66", "#3300DD"), bias=0.6)(nCol)
# lets set the information scores from single informations to grey. We
# change the highest level of the spectrum to grey.
#spect[nCol] <- "#CCCCCC"
Icol <- vector()
for (i in 1:length(I)) {
Icol[i] <- spect[ IP[i] ]
}
plot(1,1, xlim=c(0, lAli), ylim=c(-0.5, 5) ,
type="n", bty="n", xlab="position in alignment", ylab="Information (bits)")
# plot as rectangles: height is information and color is coded to information
for (i in 1:lAli) {
rect(i, 0, i+1, I[i], border=NA, col=Icol[i])
}
# As you can see, some of the columns reach very high values, but they are not
# contiguous in sequence. Are they contiguous in structure? We will find out in
# a later assignment, when we map computed values to structure.
Links and resources
| Altenhoff & Dessimoz (2012) Inferring orthology and paralogy. Methods Mol Biol 855:259-79. (pmid: 22407712) |
|
[ PubMed ] [ DOI ] The distinction between orthologs and paralogs, genes that started diverging by speciation versus duplication, is relevant in a wide range of contexts, most notably phylogenetic tree inference and protein function annotation. In this chapter, we provide an overview of the methods used to infer orthology and paralogy. We survey both graph-based approaches (and their various grouping strategies) and tree-based approaches, which solve the more general problem of gene/species tree reconciliation. We discuss conceptual differences among the various orthology inference methods and databases, and examine the difficult issue of verifying and benchmarking orthology predictions. Finally, we review typical applications of orthologous genes, groups, and reconciled trees and conclude with thoughts on future methodological developments. |
Footnotes and references
Ask, if things don't work for you!
- If anything about the assignment is not clear to you, please ask on the mailing list. You can be certain that others will have had similar problems. Success comes from joining the conversation.
- Do consider how to ask your questions so that a meaningful answer is possible:
- How to create a Minimal, Complete, and Verifiable example on stackoverflow and ...
- How to make a great R reproducible example are required reading.
