UCSF Chimera
UCSF Chimera
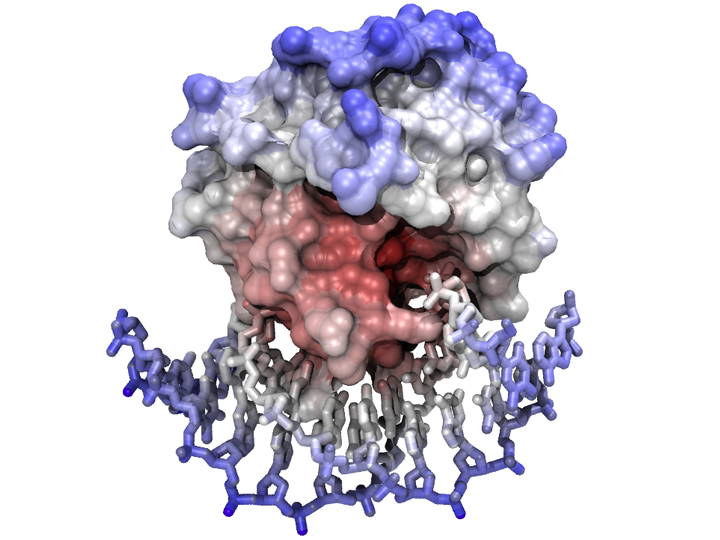
The model was visualized and positioned in VMD - nucleotides were displayed as "licorice" sticks and the protein was covered by its solvent-accessible surface, calculated with a probe radius of 1.4 Å. Coloring is ramped according to position (i.e. an atom's distance from the complex's centre of mass). The scene was rendered with the tachyon raytracer using ambient occlusion lighting and 8-fold antialiasing.
UCSF Chimera is a powerful, well engineered, molecular graphics package with a large number of functions that is under ongoing development and maintenance. It is one of a number of freely available tools to visualize and manipulate three-dimensional molecular datasets, but it stands out due to its rich features, the versatility of its use and the ease with which common tasks can be accomplished and the fact that it is being actively developed and maintained. You should learn to work with it in this course, but you should also be aware that there are alternatives, each with different strengths and weaknesses. Some of the more widely used free systems are:
A comprehensive list of molecular graphics systems can be found here.
Installing Chimera
Task:
- Access the Chimera homepage and navigate to the Download section.
- Follow the the newest version for your platform in the table and click on the file to download it.
- Follow the instructions to install Chimera.
Chimera tutorials
The Chimera User Guide site has a set of associated tutorials and there are more tutorials linked from here.