BioCyc
BioCyc.org.jpg
http://biocyc.org/META/NEW-IMAGE?type=PATHWAY&object=PWY-6805&detail-level=2
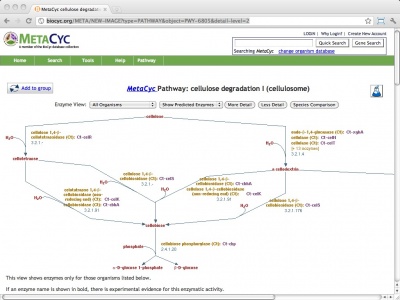
BioCyc is a collection of metabolic pathways databases, derived from computational annotation (and manual curation) of whole-genome sequence data. The database range from highly curated (such as EcoCyc, and HumanCyc, and the comparative, multiorganism MetaCyc resource) to purely computationally derived. Searches can be performed by component, reaction or pathway, and by ontology. The example URL leads to the cellulosome cellulose degradation pathway in MetaCyc.
===Reference===
| Latendresse et al. (2012) Browsing metabolic and regulatory networks with BioCyc. Methods Mol Biol 804:197-216. (pmid: 22144155) |
|
[ PubMed ] [ DOI ] AbstractThe BioCyc database collection at BioCyc.org integrates genome and cellular network information for more than 1,100 organisms. This method chapter describes Web-based tools for browsing metabolic and regulatory networks within BioCyc. These tools allow visualization of complete metabolic and regulatory networks, and allow the user to zoom-in on regions of the network of interest. The user can find objects of interest such as genes and metabolites within the networks, and can selectively examine the connectivity of the network. The EcoCyc database within the BioCyc collection has been extensively curated. The descriptions within EcoCyc of the Escherichia coli metabolic network and regulatory network were derived from thousands of publications. Other BioCyc databases received moderate levels of curation, or no curation at all. Those databases receiving no curation contain metabolic networks that were computationally inferred from the annotated genome sequences of each organism.
|
URL
http://biocyc.org/META/NEW-IMAGE?type=PATHWAY&object=PWY-6805&detail-level=2
Abstract
BioCyc is a collection of metabolic pathways databases, derived from computational annotation (and manual curation) of whole-genome sequence data. The database range from highly curated (such as EcoCyc, and HumanCyc, and the comparative, multiorganism MetaCyc resource) to purely computationally derived. Searches can be performed by component, reaction or pathway, and by ontology. The example URL leads to the cellulosome cellulose degradation pathway in MetaCyc.
Reference
| Latendresse et al. (2012) Browsing metabolic and regulatory networks with BioCyc. Methods Mol Biol 804:197-216. (pmid: 22144155) |
|
[ PubMed ] [ DOI ] AbstractThe BioCyc database collection at BioCyc.org integrates genome and cellular network information for more than 1,100 organisms. This method chapter describes Web-based tools for browsing metabolic and regulatory networks within BioCyc. These tools allow visualization of complete metabolic and regulatory networks, and allow the user to zoom-in on regions of the network of interest. The user can find objects of interest such as genes and metabolites within the networks, and can selectively examine the connectivity of the network. The EcoCyc database within the BioCyc collection has been extensively curated. The descriptions within EcoCyc of the Escherichia coli metabolic network and regulatory network were derived from thousands of publications. Other BioCyc databases received moderate levels of curation, or no curation at all. Those databases receiving no curation contain metabolic networks that were computationally inferred from the annotated genome sequences of each organism.
|