Difference between revisions of "BIO Assignment 4 2011"
m (moved Assignment 4 to BIO Assignment 4 2011) |
|||
| (10 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
| − | < | + | <!-- {{Template:Inactive}} --> |
| − | + | {{Template:Active}} | |
| − | |||
| − | |||
| − | |||
| − | |||
| Line 13: | Line 9: | ||
<div style="padding: 5px; background: #A6AFD0; border:solid 1px #AAAAAA; font-size:200%;font-weight:bold;"> | <div style="padding: 5px; background: #A6AFD0; border:solid 1px #AAAAAA; font-size:200%;font-weight:bold;"> | ||
| − | Assignment 4 - | + | Assignment 4 (last: 2011) - Phylogenetic Analysis |
</div> | </div> | ||
| − | + | <div style="padding: 2px; background: #F0F1F7; border:solid 1px #AAAAAA; font-size:125%;color:#444444"> | |
| − | + | Introduction | |
| − | |||
| − | |||
| − | <div style="padding: | ||
| − | |||
| − | |||
| − | |||
| | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | ;Nothing in Biology makes sense except in the light of evolution. | |
| − | + | :''Theodosius Dobzhansky'' | |
| − | |||
| − | |||
| − | |||
| − | : | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
</div> | </div> | ||
| − | + | ... but does evolution make sense in the light of biology? | |
| − | + | As we have seen in the previous assignments, the Mbp1 transcription factor has homologues in all other fungi, yet there is not always a clear one-to-one mapping between members of a family in distantly related species. It appears that various systems of APSES domain transcription factors have evolved independently. Of course this bears directly on our notion of function - what it means to say that two genes in different organisms have the "same" function. In case two organisms both have an orthologous gene for the same, distinct function, saying that the function is the same may be warranted. But what if that gene has duplicated in one species, and the two paralogues now perform different, related functions in one organism? Theses two are still orthologues to the other species, but now we expect functionally significant residues to have adapted to the new role of one paralogue. In order to be able to even ask such questions, we need to make the evolutionary history of gene families explicit. This is the domain of '''phylogenetic analysis'''. We can ask questions like: how many paralogues did the cenancestor of a clade possess? Which of these underwent additional duplications in the phylogenesis of the organism I am studying? Did any genes get lost? And - adding additional biological insight to the picture - did the observed duplications lead to the "invention" of new biological systems? When was that? And perhaps even: how did the species benefit from this event? | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | We will develop this kind of analysis in this assignment. In the previous assignment you have established which gene in your species is the reciprocally most closely related orthologue to yeast Mbp1 and you have identified the full complement of APSES domain genes in your assigned organism. In this assignment, we will analyse these genes' evolutionary relationship and compare it to the evolutionary relationship of all fungal APSES domains. The goal is to define families of related transcription factors and their evolutionary history. | |
| − | + | A number of excellent tools for phylogenetic analysis exist; ''general purpose packages'' include the (free) [http://evolution.genetics.washington.edu/phylip.html PHYLIP] package and the (commercial) PAUP package. ''Specialized tools'' for tree-building include Treepuzzle or Mr. Bayes. This assignment is constructed around programs that are available in PHYLIP, however you are welcome to use other tools that fulfill a similar purpose if you wish. In this field, researchers consider trees that have been built with ML (maximum likelihood) methods to be more reliable than trees that are built with parsimony methods, or distance methods such as NJ (Neighbor Joining). However ML methods are also much more compute-intensive. Just like with multiple sequence alignments, some algorithms will come closer to guessing the truth and others will not and usually it is hard to tell which is the more trustworthy of two diverging results. The prudent researcher tries out alternatives and forms her own opinion. Specifically, we may usually assume results that converge when computed with different algorithms, to be more reliable than those that depend strongly on a particular algorithm, parameters, or details of input data. | |
| − | |||
| − | |||
| − | + | However: regarding algorithm and resources, we will take a shortcut in this assignment (something you should not do in real life). We will assume that the tree the algorithm constructs is correct. In "real life" you would establish its reliability with a bootstrap procedure: repeat the tree-building a hundred times with partial data and see which branches and groupings are robust and which depend on the details of the data. In this assignment, we should simply acknowledge that bifurcations that are very close to each other have not been "resolved" and be appropriately cautious in our inferences. In phylogenetic analysis, not all lines a program draws are equally trustworthy. Don't take the trees as a given fact just because a program suggests this. Look at the evidence, include independent information where available, use your reasoning, and analyse the results critically. As you will see, there are some facts that we know for certain: we know which species the genes we have sequenced come from, and we can (usually) make good assumptions about the relationship of the species themselves - the history of speciation events that underlies all evolution of genes. This is extremely helpful information for our work. | |
| − | + | =====Introduction: Tasks===== | |
| − | + | For this assignment, we start from the APSES domains you have collected previously. You will align these domains with a set of reference domains and edit the alignment to make it suitable for phylogenetic analysis, using Jalview. Then you will construct a phylogenetic tree and interpret the tree. The goal is to identify orthologues and paralogues. <!-- Optionally, you will look at structural and functional conservation of residues. --> | |
| − | + | In case you want to review concept of trees, clades, LCAs, OTUs and the like, I have linked an excellent and very understandable introduction-level [http://biochemistry.utoronto.ca/undergraduates/courses/BCH441H/restricted/Baldauf_2003_PhylogenyTutorial.pdf article on phylogenetic analysis (pdf)] here and to the resource section at the bottom of this page. | |
| − | + | | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | {{Template:Preparation| | |
| + | care=Be sure you have understood all parts of the assignment and cover all questions in your answers! Sadly, we always get assignments back in which important aspects have simply overlooked marks unnecessarily. If you did not notice that the above did not make sense, you are reading what you expect, not what is written.| | ||
| + | num=4| | ||
| + | ord=fourth| | ||
| + | due = Monday, November 28 at 12:00 in the morning}} | ||
| − | + | ;Your documentation for the procedures you follow in this assignment will be worth 1 mark. | |
| − | |||
| − | |||
| | ||
| Line 100: | Line 51: | ||
<div style="padding: 5px; background: #BDC3DC; border:solid 1px #AAAAAA;"> | <div style="padding: 5px; background: #BDC3DC; border:solid 1px #AAAAAA;"> | ||
| − | ==(1) | + | |
| + | ==(1) Preparations== | ||
</div> | </div> | ||
| − | + | | |
| − | + | | |
| − | |||
<div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | <div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | ||
| − | === | + | ===(1.1) Preparing Input Files=== |
</div> | </div> | ||
<br> | <br> | ||
| − | |||
| − | + | For this assignment, we start from the multiple sequence alignments we have constructed previously. We will edit the alignment to make it suitable for phylogenetic analysis. We will construct a phylogenetic tree and we will analyse the tree. | |
| − | + | =====Introduction: Principle===== | |
| − | : | + | In order to use molecular sequences for the construction of phylogenetic trees, you have to build a multiple alignment first, then edit it. This is important: all rows of sequences have to contain the exact same number of characters and to hold '''aligned characters in corresponding positions'''. Phylogeny programs are not meant to revise an alignment but to analyze evolutionary relationships, '''after''' the alignment has been determined. The program's inferences are made on a column-wise basis and if your columns contain data from unrelated positions, the inferences are going to be questionable. |
| − | |||
| − | |||
| − | + | The result of the tree construction is a decision about the most likely evolutionary relationships. Fundamentally, tree-construction programs decide which sequences had common ancestors. | |
| − | |||
| − | + | '''Distance based''' phylogeny programs start by using sequence comparisons to estimate evolutionary distances: | |
| + | * they apply a model of evolution such as a mutation data matrix, to calculate a score for each pair of sequences, | ||
| + | * this score is stored in a "distance matrix" ... | ||
| + | * ... and used to estimate a tree that groups sequences with close relationships together. (e.g. by using an NJ, Neigbor Joining, algorithm). | ||
| − | + | They are fast, can work on large numbers of sequences, but are less accurate if genes evolve at different rates. | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | '''Parsimony based''' phylogeny programs build a tree that minimizes the number of mutation events that are required to get from a common ancestral sequence to all observed sequences. They take all columns into account, not just a single number per sequence pair, as the Distance Methods do. For closely related sequences they work very well, but they construct inaccurate trees when they can't make good estimates for the required number of sequence changes. | |
| − | |||
| − | + | '''ML''', or '''Maximum Likelihood''' methods attempt to find the tree for which the observed sequences would be the most likely under a particular evolutionary model. They are based on a rigorous statistical framework and yield the most robust results. But they are also quite compute intensive and a tree of the size that we are building in this assignment is a challenge for the resources of common workstation (runs about an hour on my computer). If the problem is too large, one may split a large problem into smaller, obvious subtrees (e.g. analysing orthologues as a group, only including a few paralogues for comparison) and then merge the smaller trees; this way even very large problems can become tractable. | |
| − | + | ML methods suffer less from "long-branch attraction" - the phenomenon that weakly similar sequences can be grouped inappropriately close together in a tree due to spuriously shared differences. | |
| − | |||
| − | + | Clearly, in order for tree-estimation to work, one must not include fragments of sequence which have evolved under a different evolutionary model as all others, e.g. after domain fusion, or after accommodating large stretches of indels. Thus it is appropriate to edit the sequences and pare them down to a most characteristic subset of amino acids. The goal is not to be as comprehensive as possible, but to input those columns of aligned residues that will best represent the true phylogenetic relationships between the sequences. | |
| − | + | =====Introduction: Gaps===== | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | Gaps are a real problem here, as usual. Strictly speaking, the similarity score of an '''alignment''' program as well as the distance score of a '''phylogeny''' program are not calculated for an ordered sequence, but for a sum of independent values, one for each aligned columns of characters. The order of the columns does not change the score. However in an optimal sequence alignment with gaps, this is no longer strictly true since a one-character gap creation has a different penalty score than a one-character gap extension! Most '''alignment''' programs use a model with a constant gap insertion penalty and a linear gap extension penalty. This is not rigorously justified from biology, but parametrized (or you could say "tweaked") to correspond to our observations. However, most '''phylogeny''' programs, (such as the programs in PHYLIP) do not work in this way. PHYLIP strictly operates on columns of characters and treats a gap character just like a residue with the one letter code "-". Thus gap insertion- and extension- characters get the same score. For short indels, this '''underestimates''' the distance between pairs of sequences, since any evolutionary model should reflect the fact that gaps are much less likely than point mutations. If the gap is very long though, all events are counted individually as many single substitutions (rather than one lengthy one) and this '''overestimates''' the distance. And it gets worse: long stretches of gaps can make sequences appear similar in a way that is not justified, just because they are identical in the "-" character. It is therefore common and acceptable to edit gaps in the alignment and delete all but one or two columns of gapped sequence, or to remove such columns altogether. | |
| − | |||
| − | + | =====Introduction: The outgroup===== | |
| − | + | To analyse phylogenetic trees it is useful (and for some algorithms required) to define an outgroup, a sequence that presumably diverged from all other sequences in a clade before they split up among themselves. Wherever the outgroup inserts into the tree, this is the root of the rest of the tree. And whenever a molecular clock is assumed, the branching point that connects the outgroup can be assumed to be the oldest divergence event. I have defined an outgroup sequence and added it to the [[Reference APSES domains|reference APSES domains page]]. The procedure is explained in detail on that page. | |
| + | >gi|301025594|ref|ZP_07189117.1| KilA-N domain protein [Escherichia coli MS 69-1] | ||
| + | <span style="color: #999999;">MTSFQLSLISRE</span>IDGEIIHLRAKDGYINATSMCRTAGKLLSDYTRLKTTQEFFDELSRDMGIPISELIQS | ||
| + | FKGGRPENQGTWVHPDIAINLAQ<span style="color: #999999;">WLSPKFAVQVSRWVREWMSGERTTAEMPVHLKRYMVNRSRIPHTHFS | ||
| + | ILNELTFNLVAPLEQAGYTLPEKMVPDISQGRVFSQWLRDNRNVEPKTFPTYDHEYPDGRVYPARLYPNE | ||
| + | YLADFKEHFNNIWLPQYAPKYFADRDKKALALIEKIMLPNLDGNEQF</span> | ||
| + | ''E. coli'' KilA-N protein. Residues that do not align with APSES domains are shown in grey. | ||
| − | + | =====Preparing APSES sequences===== | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | < | + | <div style="padding: 5px; background: #DDDDEE;"> |
| − | + | #Navigate to the [[Reference APSES domains|reference APSES domains page]] and copy the sequences. | |
| − | + | #Open Jalview, select '''File → Input Alignment → from Textbox''' and paste the sequences into the textbox. | |
| − | </ | + | #Add the APSES domain sequences '''from your species''' that you have defined in the previous assignment. |
| + | #When all the sequences are present, click on '''New Window'''. | ||
| + | #In Jalview, select Web Service → Alignment → MAFFT Multiple Sequence Alignment. The alignment is calculated in a few minutes and displayed in a new window. | ||
| + | #Choose any colour scheme and add '''Colour → by Conservation'''. Adjust the slider left or right to see which columns are highly conserved. | ||
| + | #Save the alignment as a Jalview project before editing it for phylogenetic analysis. You may need it again. | ||
| + | </div> | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | =====Introduction: Alignment editing for phylogenetic reconstruction===== | |
| − | |||
| − | |||
| − | |||
| − | + | In practice, follow the fundamental principle that '''all characters in a column should be related by homology'''. This implies the following rules of thumb: | |
| − | |||
| − | |||
| − | |||
| − | + | *Remove all stretches of residues in which the ''alignment'' appears ambiguous (not just highly variable, but ambiguous regarding the aligned positions). | |
| − | + | *Remove all frayed N- and C- termini, especially regions in which not all sequences that are being compared appear homologous and that may stem from unrelated domains. | |
| − | + | *Remove all gapped regions that appear to be alignment artefacts due to inappropriate input sequences. | |
| − | + | *Remove all but approximately one column from gapped regions '''in those cases where the presence of several related insertions suggest that the indel is real, and not just an alignment artefact.''' (Some researchers simply remove all gapped regions). | |
| + | *Remove sections N- and C- terminal of gaps where the alignment appears questionable. | ||
| + | *Also, consider that neither residues that are completely different between all species, nor residues that are completely conserved are informative for relationship distances. | ||
| + | *If your sequences are too long, your tree calculations may run out of memory. 60-80 aligned residues should be plenty and if the sequences fit on a single line you will save yourself potential trouble with block-wise vs. interleaved input.If you do run out of memory try removing columns of sequence. | ||
| + | *Move the KilA-N outgroup sequence to the first line of your alignment, since this is where PHYLIP will look for it by default. | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | [[Image:EditingGuide.jpg|frame|none|(Possible) steps in editing a multiple sequence alignment towards a PHYLIP input file. '''a''': raw alignment (CLUSTAL format); '''b''': sequences assembled into single lines; '''c''': columns to be deleted highlighted in red - 1, 3 and 4: large gaps; 2: uncertain alignment and 5: frayed C-terminus: both would put non-homologous characters into the same column; '''d''': input data for PHYLIP: names for sequences must not be longer than 10 characters, the first line must contain the number of sequences and the sequence length. PHYLIP is very picky about incorrectly formatted input, read the [http://evolution.genetics.washington.edu/phylip/doc/sequence.html PHYLIP sequence format guide].]] | |
| − | |||
| − | |||
| − | |||
| − | + | ;Once you are satisfied with your editing, proceed as follows: | |
| − | |||
| − | |||
| − | |||
| − | < | + | <div style="padding: 5px; background: #DDDDEE;"> |
| − | + | #Download the PHYLIP package from the [http://evolution.genetics.washington.edu/phylip.html Phylip homepage] and install it on your computer. | |
| − | + | #Prepare a PHYLIP input file from your Jalview alignment. The simplest way to achieve this appears to be: | |
| − | </ | + | ##In Jalview, use '''File → Output to Textbox→FASTA''', then '''Edit→Select All''' and '''Edit→copy''' the sequences. |
| + | ##In a browser, navigate to the [http://www-bimas.cit.nih.gov/molbio/readseq/ '''Readseq sequence conversion service''']. | ||
| + | ##Paste your sequences into the form and choose '''Phylip''' as the output format. Click on '''submit'''. | ||
| + | ##Save the resulting page as a text file in the directory where the phylip executables reside on your computer. Give it some useful name such as <code>All-APSES_domains.phy</code>. | ||
| + | #Make a copy of that file and name it <code>infile</code>. Note: make sure that your Microsoft Windows operating system does not silently append the extension ".txt" to your file. It should be called "infile", nothing else and you should never, never, ever permit your operating systems to slyly hide file extensions from you when it displays filenames. You have been warned. | ||
| + | </div> | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | < | + | <div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> |
| − | |||
| − | |||
| − | |||
| − | + | ===(1.2) Calculating a Tree=== | |
| − | + | </div> | |
| − | |||
| − | </ | ||
| − | < | + | <br> |
| − | + | <br> | |
| − | + | <div style="padding: 5px; background: #DDDDEE;"> | |
| − | |||
| − | < | + | *Use the '''proml''' program of PHYLIP (protein sequences, maximum likelihood tree) to calculate a phylogenetic tree. Use the default parameters except that you must change option <code>S: Speedier but rougher analysis?</code> to No - your analysis should not sacrifice accuracy for speed. The calculation will take a while. |
| − | |||
| − | |||
| − | |||
| − | + | </div> | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
<br> | <br> | ||
| − | + | <br> | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | | ||
<div style="padding: 5px; background: #BDC3DC; border:solid 1px #AAAAAA;"> | <div style="padding: 5px; background: #BDC3DC; border:solid 1px #AAAAAA;"> | ||
| − | ==(2) | + | ==(2) Analysis (2 marks)== |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
</div> | </div> | ||
| − | |||
| − | + | I have constructed a cladogram for the species we are analysing, based on data published for 1551 fungal ribosomal sequences. Such reference tres from rRNA data are a standard method of phylogenetic analysis, supported by the assumption that rRNA sequences are monophyletic and have evolved under comparable selective pressure in all species. | |
| − | + | [[Image:FungiCladogram.jpg|frame|none|Cladogram of fungi studied in the assignments. This cladogram is based on small subunit ribosomal rRNA sequences, and largely follows ''Tehler et al.'' (2003) ''Mycol Res.'' '''107''':901-916. Even though many details of fungal phylogeny remain unresolved, the branches shown here individually appear to have strong support. In a cladogram such as this, the branch lengths are not drawn to any scale of similarity.]] | |
| − | |||
| − | : | ||
| − | + | Your species may not be included in this cladogram, but you can easily calculate your own with the following procedure: | |
| − | + | <div style="padding: 5px; background: #DDDDEE;"> | |
| + | #Access the [http://www.ncbi.nlm.nih.gov/sites/entrez?cmd=retrieve&db=taxonomy NCBI taxonomy database Entrez query page]. | ||
| + | #Edit the list of reference species below to include your species and paste it into the form. | ||
| − | + | "Emericella nidulans"[Scientific Name] OR | |
| + | "Candida albicans"[Scientific Name] OR | ||
| + | "Neurospora crassa"[Scientific Name] OR | ||
| + | "Saccharomyces cerevisiae"[Scientific Name] OR | ||
| + | "Schizosaccharomyces pombe"[Scientific Name] OR | ||
| + | "Ustilago maydis"[Scientific Name] | ||
| + | #Next, as '''Display''' option, select '''Common Tree'''. | ||
| + | #Then select the '''phylip tree''' option and click '''save as''' to save the tree in Newick format. | ||
| + | #The output can be edited, and visualized in any program that reads Newick trees. | ||
</div> | </div> | ||
| − | |||
| − | |||
| − | <div style="padding: 5px; background: # | + | <div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> |
| − | ==( | + | ===(2.2) Visualizing the APSES domain Phylogenetic Tree=== |
</div> | </div> | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | Once Phylip is done calculating the tree, the tree in a text format will be contained in the Phylip <code>outfile</code> - the documentation of what the program has done. Open this textfile for a first look. The tree is complicated and it can look confusing at first. The tree in Newick format is contained in the Phylip file <code>outtree</code>. Visualize it as follows: | |
| − | + | <div style="padding: 5px; background: #DDDDEE;"> | |
| − | + | #Open <code>outtree</code> in a texteditor and copy the tree. | |
| + | #Visualize the tree in alternative representations: | ||
| + | ##Navigate to the [http://www.proweb.org/treeviewer/ Proweb treeviewer], paste and visualize your tree. | ||
| + | ##Navigate to the [http://www.trex.uqam.ca/index.php?action=newick&project=trex Trex-online Newick tree viewer] for an alternative view. Visualize the tree as a phylogram. You can increase the window height to keep the labels from overlapping. | ||
| + | ##In your Jalview window, choose '''File → Load associated Tree''' and load the Phylip <code>outtree</code> file. You can click into the tree-window to show which clades branch off at what level - it should be obvious that you can identify three major subclades. | ||
| + | ##Study the tree: understand what you see and what you would have expected. | ||
</div> | </div> | ||
| − | + | Here are two principles that will help you make sense of the tree. | |
| − | |||
| − | |||
| − | + | A: '''A gene that is present in an ancestral species is inherited in all descendant species'''. The gene has to be observed in all OTUs, unless its has been lost (which is a rare event). | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | B: '''Paralogous genes in an ancestral species should give rise to monophyletic subtrees for each of the paralogues, in all descendants'''; this means: if the LCA of a branch has e.g. three genes, we would expect three copies of the species cladogram below this branchpoint, one for each of these genes. Each of these subtrees should recapitulate the reference phylogenetic tree of the species, up to the branchpoint of their LCA. | |
| − | |||
| − | |||
| − | |||
| + | With these two simple principles (you should draw them out on a piece of paper if they do not seem obvious to you), you can probably pry your tree apart quite nicely. A few colored pencils and a printout of the tree will help. | ||
| − | |||
| | ||
| Line 369: | Line 227: | ||
<div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | <div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | ||
| − | ===( | + | ===(2.1) The Cenancestor's APSES Domains=== |
</div> | </div> | ||
| − | |||
| − | + | Refer to your tree for the following tasks. (Please remember to include your tree in your Assignment submission - it is a result of your computational experiment. Its easiest to copy/paste the tree from the Phylip outfile, rather than copying an image from a Tree viewer). Be specific in your discussion, i.e. refer to specific branchpoints (branchpoints are numbered in the Phylip output) and OTU or gene names in your analysis (see the example below). | |
| − | |||
| − | + | <div style="padding: 5px; background: #DDDDEE;"> | |
| + | *Consider how many APSES domain proteins the fungal cenancestor appears to have possessed and what evidence you see in the tree that this is so. | ||
| + | </div> | ||
| − | |||
| − | + | <div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | |
| − | + | ===(2.2) Unraveling your organism's APSES domains (2 marks)=== | |
| + | </div> | ||
| − | + | <br> | |
| + | <br> | ||
| + | <div style="padding: 5px; background: #FFCC99;"> | ||
| + | ;Analysis (2 marks) | ||
| − | + | Assume that the cladogram for fungi that I have given above is correct, and that the mixed gene tree you have calculated is fundamentally correct in its overall arrangement but may have local inaccuracies due to the limited resolution of the method. You have identified the APSES domain genes of the fungal cenancestor above. Apply the expectations we have stated above to discuss briefly through what sequence of duplications and/or gene loss your organism has ended up with the APSES domains it possesses today. Make specific reference to the cladogram of species and note in particular in case some of your sequences appear to have been placed into regions of the tree where they don't seem to belong. Also note which branchpoints in the evolutionary history of your sequences correspond to speciations and which ones to duplications. | |
| − | |||
| − | + | Note: A common confusion about cenancestral genes arises from the fact that by far not all expected genes are present in the OTUs. Some will have been lost, some will have been incorrectly annotated in their genome (frameshifts!) and not been found with PSI-BLAST, some may have been missed by you. In general you have to ask: '''given the species represented in a subclade, what is the last common ancestor of that branch'''? The expectation is that '''all''' descendants of that ancestor should be represented in that branch '''unless''' one of the above reasons why a gene might be absent would apply. | |
| − | |||
| − | + | If your species does not have all the genes you would expect it to have inherited from its ancestors, you MUST note that fact and attempt to explain it. | |
| − | + | </div> | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | For example the following discusion for ''Saccharomyces cerevisiae'' would be sufficient for full marks: | |
| − | + | :(Numbers refer to branchpoints of the mixed gene tree, letters to branchpoints of the species tree). I have found five homologues to ''Saccharomyces cerevisiae'' Mbp1 and included them in the mixed gene tree. Two subclades are well defined, and contain all current species, they branch from 41 (Xbp1) and 50 (Sok2/Phd1). The subclade below 6 includes Mbp1 orthologues as well as Swi4 orthologues that do not appear well resolved. Considering only species below the ''saccharomycetales'' branchpoint, I postulate a duplication at that branchpoint that gave rise to yeast Mbp1 and Swi4 since the respective branches contain representatives from all fungi that descended from that branch. There is no good support for the idea that the cenancestor had a Swi4 paralogue. Therefore the cenancestor most likely posessed two paralogues: Mbp1, and Sok2. ''Saccharomyces cerevisiae'' has one gene in each of the major subclades, there is no gene loss. It also has an additional paralogue to Sok2: the Phd1 gene that duplicated at branchpoint 3. | |
| − | |||
| − | |||
| − | |||
| − | |||
<div style="padding: 5px; background: #BDC3DC; border:solid 1px #AAAAAA;"> | <div style="padding: 5px; background: #BDC3DC; border:solid 1px #AAAAAA;"> | ||
| − | ==( | + | ==(3) Summary of Resources== |
</div> | </div> | ||
<br> | <br> | ||
;Links | ;Links | ||
| − | :* [http://biochemistry.utoronto.ca/undergraduates/courses/BCH441H/restricted/ | + | :* [http://biochemistry.utoronto.ca/undergraduates/courses/BCH441H/restricted/Baldauf_2003_PhylogenyTutorial.pdf '''Review (PDF, restricted)''' Sandra Baldauf: Phylogeny for the Faint of Heart] |
| − | : | + | :* [http://evolution.genetics.washington.edu/phylip.html '''PHYLIP''' home page] |
| − | |||
| − | :* [http:// | ||
| − | |||
| − | + | ;Sequences | |
| − | + | :* [[Reference APSES domains|Reference APSES domains page]] | |
| − | ; | ||
| − | :* [[ | ||
| − | |||
| − | |||
| − | |||
<div style="padding: 5px; background: #D3D8E8; border:solid 1px #AAAAAA;"> | <div style="padding: 5px; background: #D3D8E8; border:solid 1px #AAAAAA;"> | ||
| Line 441: | Line 279: | ||
</div> | </div> | ||
| − | If you have any questions at all, don't hesitate to mail me at [mailto:boris.steipe@utoronto.ca boris.steipe@utoronto.ca] or post your question to the [mailto: | + | If you have any questions at all, don't hesitate to mail me at [mailto:boris.steipe@utoronto.ca boris.steipe@utoronto.ca] or post your question to the [mailto:bch441_2011@googlegroups.com Course Mailing List] |
Latest revision as of 23:34, 21 September 2012
Note! This assignment is currently active. All significant changes will be announced on the mailing list.
Contents
Assignment 4 (last: 2011) - Phylogenetic Analysis
Introduction
- Nothing in Biology makes sense except in the light of evolution.
- Theodosius Dobzhansky
... but does evolution make sense in the light of biology?
As we have seen in the previous assignments, the Mbp1 transcription factor has homologues in all other fungi, yet there is not always a clear one-to-one mapping between members of a family in distantly related species. It appears that various systems of APSES domain transcription factors have evolved independently. Of course this bears directly on our notion of function - what it means to say that two genes in different organisms have the "same" function. In case two organisms both have an orthologous gene for the same, distinct function, saying that the function is the same may be warranted. But what if that gene has duplicated in one species, and the two paralogues now perform different, related functions in one organism? Theses two are still orthologues to the other species, but now we expect functionally significant residues to have adapted to the new role of one paralogue. In order to be able to even ask such questions, we need to make the evolutionary history of gene families explicit. This is the domain of phylogenetic analysis. We can ask questions like: how many paralogues did the cenancestor of a clade possess? Which of these underwent additional duplications in the phylogenesis of the organism I am studying? Did any genes get lost? And - adding additional biological insight to the picture - did the observed duplications lead to the "invention" of new biological systems? When was that? And perhaps even: how did the species benefit from this event?
We will develop this kind of analysis in this assignment. In the previous assignment you have established which gene in your species is the reciprocally most closely related orthologue to yeast Mbp1 and you have identified the full complement of APSES domain genes in your assigned organism. In this assignment, we will analyse these genes' evolutionary relationship and compare it to the evolutionary relationship of all fungal APSES domains. The goal is to define families of related transcription factors and their evolutionary history.
A number of excellent tools for phylogenetic analysis exist; general purpose packages include the (free) PHYLIP package and the (commercial) PAUP package. Specialized tools for tree-building include Treepuzzle or Mr. Bayes. This assignment is constructed around programs that are available in PHYLIP, however you are welcome to use other tools that fulfill a similar purpose if you wish. In this field, researchers consider trees that have been built with ML (maximum likelihood) methods to be more reliable than trees that are built with parsimony methods, or distance methods such as NJ (Neighbor Joining). However ML methods are also much more compute-intensive. Just like with multiple sequence alignments, some algorithms will come closer to guessing the truth and others will not and usually it is hard to tell which is the more trustworthy of two diverging results. The prudent researcher tries out alternatives and forms her own opinion. Specifically, we may usually assume results that converge when computed with different algorithms, to be more reliable than those that depend strongly on a particular algorithm, parameters, or details of input data.
However: regarding algorithm and resources, we will take a shortcut in this assignment (something you should not do in real life). We will assume that the tree the algorithm constructs is correct. In "real life" you would establish its reliability with a bootstrap procedure: repeat the tree-building a hundred times with partial data and see which branches and groupings are robust and which depend on the details of the data. In this assignment, we should simply acknowledge that bifurcations that are very close to each other have not been "resolved" and be appropriately cautious in our inferences. In phylogenetic analysis, not all lines a program draws are equally trustworthy. Don't take the trees as a given fact just because a program suggests this. Look at the evidence, include independent information where available, use your reasoning, and analyse the results critically. As you will see, there are some facts that we know for certain: we know which species the genes we have sequenced come from, and we can (usually) make good assumptions about the relationship of the species themselves - the history of speciation events that underlies all evolution of genes. This is extremely helpful information for our work.
Introduction: Tasks
For this assignment, we start from the APSES domains you have collected previously. You will align these domains with a set of reference domains and edit the alignment to make it suitable for phylogenetic analysis, using Jalview. Then you will construct a phylogenetic tree and interpret the tree. The goal is to identify orthologues and paralogues.
In case you want to review concept of trees, clades, LCAs, OTUs and the like, I have linked an excellent and very understandable introduction-level article on phylogenetic analysis (pdf) here and to the resource section at the bottom of this page.
Preparation, submission and due date
- Read carefully.
- Be sure you have understood all parts of the assignment and cover all questions in your answers! Sadly, we always get assignments back in which important aspects have simply overlooked marks unnecessarily. If you did not notice that the above did not make sense, you are reading what you expect, not what is written.
Review the guidelines for preparation and submission of BCH441 assignments.
The due date for the assignment is Monday, November 28 at 12:00 in the morning.
- Your documentation for the procedures you follow in this assignment will be worth 1 mark.
(1) Preparations
(1.1) Preparing Input Files
For this assignment, we start from the multiple sequence alignments we have constructed previously. We will edit the alignment to make it suitable for phylogenetic analysis. We will construct a phylogenetic tree and we will analyse the tree.
Introduction: Principle
In order to use molecular sequences for the construction of phylogenetic trees, you have to build a multiple alignment first, then edit it. This is important: all rows of sequences have to contain the exact same number of characters and to hold aligned characters in corresponding positions. Phylogeny programs are not meant to revise an alignment but to analyze evolutionary relationships, after the alignment has been determined. The program's inferences are made on a column-wise basis and if your columns contain data from unrelated positions, the inferences are going to be questionable.
The result of the tree construction is a decision about the most likely evolutionary relationships. Fundamentally, tree-construction programs decide which sequences had common ancestors.
Distance based phylogeny programs start by using sequence comparisons to estimate evolutionary distances:
- they apply a model of evolution such as a mutation data matrix, to calculate a score for each pair of sequences,
- this score is stored in a "distance matrix" ...
- ... and used to estimate a tree that groups sequences with close relationships together. (e.g. by using an NJ, Neigbor Joining, algorithm).
They are fast, can work on large numbers of sequences, but are less accurate if genes evolve at different rates.
Parsimony based phylogeny programs build a tree that minimizes the number of mutation events that are required to get from a common ancestral sequence to all observed sequences. They take all columns into account, not just a single number per sequence pair, as the Distance Methods do. For closely related sequences they work very well, but they construct inaccurate trees when they can't make good estimates for the required number of sequence changes.
ML, or Maximum Likelihood methods attempt to find the tree for which the observed sequences would be the most likely under a particular evolutionary model. They are based on a rigorous statistical framework and yield the most robust results. But they are also quite compute intensive and a tree of the size that we are building in this assignment is a challenge for the resources of common workstation (runs about an hour on my computer). If the problem is too large, one may split a large problem into smaller, obvious subtrees (e.g. analysing orthologues as a group, only including a few paralogues for comparison) and then merge the smaller trees; this way even very large problems can become tractable.
ML methods suffer less from "long-branch attraction" - the phenomenon that weakly similar sequences can be grouped inappropriately close together in a tree due to spuriously shared differences.
Clearly, in order for tree-estimation to work, one must not include fragments of sequence which have evolved under a different evolutionary model as all others, e.g. after domain fusion, or after accommodating large stretches of indels. Thus it is appropriate to edit the sequences and pare them down to a most characteristic subset of amino acids. The goal is not to be as comprehensive as possible, but to input those columns of aligned residues that will best represent the true phylogenetic relationships between the sequences.
Introduction: Gaps
Gaps are a real problem here, as usual. Strictly speaking, the similarity score of an alignment program as well as the distance score of a phylogeny program are not calculated for an ordered sequence, but for a sum of independent values, one for each aligned columns of characters. The order of the columns does not change the score. However in an optimal sequence alignment with gaps, this is no longer strictly true since a one-character gap creation has a different penalty score than a one-character gap extension! Most alignment programs use a model with a constant gap insertion penalty and a linear gap extension penalty. This is not rigorously justified from biology, but parametrized (or you could say "tweaked") to correspond to our observations. However, most phylogeny programs, (such as the programs in PHYLIP) do not work in this way. PHYLIP strictly operates on columns of characters and treats a gap character just like a residue with the one letter code "-". Thus gap insertion- and extension- characters get the same score. For short indels, this underestimates the distance between pairs of sequences, since any evolutionary model should reflect the fact that gaps are much less likely than point mutations. If the gap is very long though, all events are counted individually as many single substitutions (rather than one lengthy one) and this overestimates the distance. And it gets worse: long stretches of gaps can make sequences appear similar in a way that is not justified, just because they are identical in the "-" character. It is therefore common and acceptable to edit gaps in the alignment and delete all but one or two columns of gapped sequence, or to remove such columns altogether.
Introduction: The outgroup
To analyse phylogenetic trees it is useful (and for some algorithms required) to define an outgroup, a sequence that presumably diverged from all other sequences in a clade before they split up among themselves. Wherever the outgroup inserts into the tree, this is the root of the rest of the tree. And whenever a molecular clock is assumed, the branching point that connects the outgroup can be assumed to be the oldest divergence event. I have defined an outgroup sequence and added it to the reference APSES domains page. The procedure is explained in detail on that page.
>gi|301025594|ref|ZP_07189117.1| KilA-N domain protein [Escherichia coli MS 69-1] MTSFQLSLISREIDGEIIHLRAKDGYINATSMCRTAGKLLSDYTRLKTTQEFFDELSRDMGIPISELIQS FKGGRPENQGTWVHPDIAINLAQWLSPKFAVQVSRWVREWMSGERTTAEMPVHLKRYMVNRSRIPHTHFS ILNELTFNLVAPLEQAGYTLPEKMVPDISQGRVFSQWLRDNRNVEPKTFPTYDHEYPDGRVYPARLYPNE YLADFKEHFNNIWLPQYAPKYFADRDKKALALIEKIMLPNLDGNEQF
E. coli KilA-N protein. Residues that do not align with APSES domains are shown in grey.
Preparing APSES sequences
- Navigate to the reference APSES domains page and copy the sequences.
- Open Jalview, select File → Input Alignment → from Textbox and paste the sequences into the textbox.
- Add the APSES domain sequences from your species that you have defined in the previous assignment.
- When all the sequences are present, click on New Window.
- In Jalview, select Web Service → Alignment → MAFFT Multiple Sequence Alignment. The alignment is calculated in a few minutes and displayed in a new window.
- Choose any colour scheme and add Colour → by Conservation. Adjust the slider left or right to see which columns are highly conserved.
- Save the alignment as a Jalview project before editing it for phylogenetic analysis. You may need it again.
Introduction: Alignment editing for phylogenetic reconstruction
In practice, follow the fundamental principle that all characters in a column should be related by homology. This implies the following rules of thumb:
- Remove all stretches of residues in which the alignment appears ambiguous (not just highly variable, but ambiguous regarding the aligned positions).
- Remove all frayed N- and C- termini, especially regions in which not all sequences that are being compared appear homologous and that may stem from unrelated domains.
- Remove all gapped regions that appear to be alignment artefacts due to inappropriate input sequences.
- Remove all but approximately one column from gapped regions in those cases where the presence of several related insertions suggest that the indel is real, and not just an alignment artefact. (Some researchers simply remove all gapped regions).
- Remove sections N- and C- terminal of gaps where the alignment appears questionable.
- Also, consider that neither residues that are completely different between all species, nor residues that are completely conserved are informative for relationship distances.
- If your sequences are too long, your tree calculations may run out of memory. 60-80 aligned residues should be plenty and if the sequences fit on a single line you will save yourself potential trouble with block-wise vs. interleaved input.If you do run out of memory try removing columns of sequence.
- Move the KilA-N outgroup sequence to the first line of your alignment, since this is where PHYLIP will look for it by default.
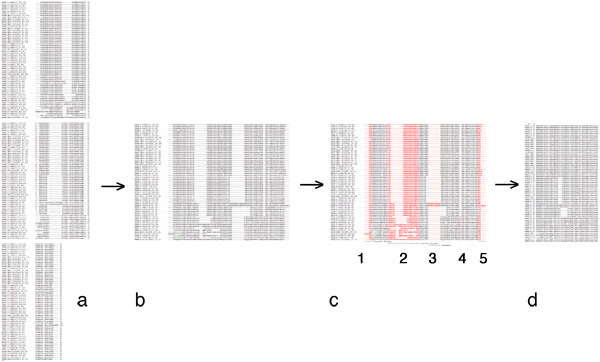

- Once you are satisfied with your editing, proceed as follows
- Download the PHYLIP package from the Phylip homepage and install it on your computer.
- Prepare a PHYLIP input file from your Jalview alignment. The simplest way to achieve this appears to be:
- In Jalview, use File → Output to Textbox→FASTA, then Edit→Select All and Edit→copy the sequences.
- In a browser, navigate to the Readseq sequence conversion service.
- Paste your sequences into the form and choose Phylip as the output format. Click on submit.
- Save the resulting page as a text file in the directory where the phylip executables reside on your computer. Give it some useful name such as
All-APSES_domains.phy.
- Make a copy of that file and name it
infile. Note: make sure that your Microsoft Windows operating system does not silently append the extension ".txt" to your file. It should be called "infile", nothing else and you should never, never, ever permit your operating systems to slyly hide file extensions from you when it displays filenames. You have been warned.
(1.2) Calculating a Tree
- Use the proml program of PHYLIP (protein sequences, maximum likelihood tree) to calculate a phylogenetic tree. Use the default parameters except that you must change option
S: Speedier but rougher analysis?to No - your analysis should not sacrifice accuracy for speed. The calculation will take a while.
(2) Analysis (2 marks)
I have constructed a cladogram for the species we are analysing, based on data published for 1551 fungal ribosomal sequences. Such reference tres from rRNA data are a standard method of phylogenetic analysis, supported by the assumption that rRNA sequences are monophyletic and have evolved under comparable selective pressure in all species.
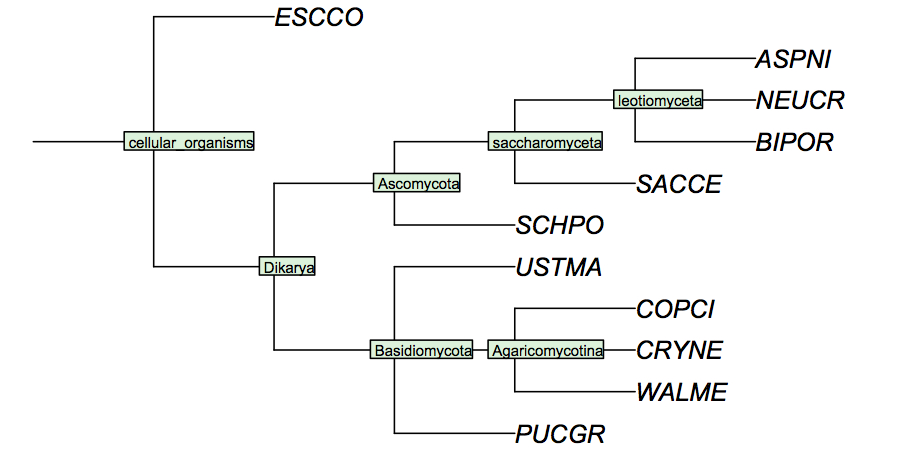
Your species may not be included in this cladogram, but you can easily calculate your own with the following procedure:
- Access the NCBI taxonomy database Entrez query page.
- Edit the list of reference species below to include your species and paste it into the form.
"Emericella nidulans"[Scientific Name] OR "Candida albicans"[Scientific Name] OR "Neurospora crassa"[Scientific Name] OR "Saccharomyces cerevisiae"[Scientific Name] OR "Schizosaccharomyces pombe"[Scientific Name] OR "Ustilago maydis"[Scientific Name]
- Next, as Display option, select Common Tree.
- Then select the phylip tree option and click save as to save the tree in Newick format.
- The output can be edited, and visualized in any program that reads Newick trees.
(2.2) Visualizing the APSES domain Phylogenetic Tree
Once Phylip is done calculating the tree, the tree in a text format will be contained in the Phylip outfile - the documentation of what the program has done. Open this textfile for a first look. The tree is complicated and it can look confusing at first. The tree in Newick format is contained in the Phylip file outtree. Visualize it as follows:
- Open
outtreein a texteditor and copy the tree. - Visualize the tree in alternative representations:
- Navigate to the Proweb treeviewer, paste and visualize your tree.
- Navigate to the Trex-online Newick tree viewer for an alternative view. Visualize the tree as a phylogram. You can increase the window height to keep the labels from overlapping.
- In your Jalview window, choose File → Load associated Tree and load the Phylip
outtreefile. You can click into the tree-window to show which clades branch off at what level - it should be obvious that you can identify three major subclades. - Study the tree: understand what you see and what you would have expected.
Here are two principles that will help you make sense of the tree.
A: A gene that is present in an ancestral species is inherited in all descendant species. The gene has to be observed in all OTUs, unless its has been lost (which is a rare event).
B: Paralogous genes in an ancestral species should give rise to monophyletic subtrees for each of the paralogues, in all descendants; this means: if the LCA of a branch has e.g. three genes, we would expect three copies of the species cladogram below this branchpoint, one for each of these genes. Each of these subtrees should recapitulate the reference phylogenetic tree of the species, up to the branchpoint of their LCA.
With these two simple principles (you should draw them out on a piece of paper if they do not seem obvious to you), you can probably pry your tree apart quite nicely. A few colored pencils and a printout of the tree will help.
(2.1) The Cenancestor's APSES Domains
Refer to your tree for the following tasks. (Please remember to include your tree in your Assignment submission - it is a result of your computational experiment. Its easiest to copy/paste the tree from the Phylip outfile, rather than copying an image from a Tree viewer). Be specific in your discussion, i.e. refer to specific branchpoints (branchpoints are numbered in the Phylip output) and OTU or gene names in your analysis (see the example below).
- Consider how many APSES domain proteins the fungal cenancestor appears to have possessed and what evidence you see in the tree that this is so.
(2.2) Unraveling your organism's APSES domains (2 marks)
- Analysis (2 marks)
Assume that the cladogram for fungi that I have given above is correct, and that the mixed gene tree you have calculated is fundamentally correct in its overall arrangement but may have local inaccuracies due to the limited resolution of the method. You have identified the APSES domain genes of the fungal cenancestor above. Apply the expectations we have stated above to discuss briefly through what sequence of duplications and/or gene loss your organism has ended up with the APSES domains it possesses today. Make specific reference to the cladogram of species and note in particular in case some of your sequences appear to have been placed into regions of the tree where they don't seem to belong. Also note which branchpoints in the evolutionary history of your sequences correspond to speciations and which ones to duplications.
Note: A common confusion about cenancestral genes arises from the fact that by far not all expected genes are present in the OTUs. Some will have been lost, some will have been incorrectly annotated in their genome (frameshifts!) and not been found with PSI-BLAST, some may have been missed by you. In general you have to ask: given the species represented in a subclade, what is the last common ancestor of that branch? The expectation is that all descendants of that ancestor should be represented in that branch unless one of the above reasons why a gene might be absent would apply.
If your species does not have all the genes you would expect it to have inherited from its ancestors, you MUST note that fact and attempt to explain it.
For example the following discusion for Saccharomyces cerevisiae would be sufficient for full marks:
- (Numbers refer to branchpoints of the mixed gene tree, letters to branchpoints of the species tree). I have found five homologues to Saccharomyces cerevisiae Mbp1 and included them in the mixed gene tree. Two subclades are well defined, and contain all current species, they branch from 41 (Xbp1) and 50 (Sok2/Phd1). The subclade below 6 includes Mbp1 orthologues as well as Swi4 orthologues that do not appear well resolved. Considering only species below the saccharomycetales branchpoint, I postulate a duplication at that branchpoint that gave rise to yeast Mbp1 and Swi4 since the respective branches contain representatives from all fungi that descended from that branch. There is no good support for the idea that the cenancestor had a Swi4 paralogue. Therefore the cenancestor most likely posessed two paralogues: Mbp1, and Sok2. Saccharomyces cerevisiae has one gene in each of the major subclades, there is no gene loss. It also has an additional paralogue to Sok2: the Phd1 gene that duplicated at branchpoint 3.
(3) Summary of Resources
- Sequences
[End of assignment]
If you have any questions at all, don't hesitate to mail me at boris.steipe@utoronto.ca or post your question to the Course Mailing List