Difference between revisions of "KEGG"
(Created page with "<div id="CSB"> <div class="b1"> KEGG </div> {{dev}} The Kyoto Encyclopedia of Genes and Genomes (KEGG) is a resource that integrates genomic, chemical and systemic functio...") |
m |
||
| Line 12: | Line 12: | ||
__TOC__ | __TOC__ | ||
| − | |||
==Contents== | ==Contents== | ||
... | ... | ||
| + | |||
| + | | ||
==Introductory reading== | ==Introductory reading== | ||
<section begin=reading /> | <section begin=reading /> | ||
| + | {{#pmid: 22080510}} <!-- intro. reading --> | ||
<section end=reading /> | <section end=reading /> | ||
| + | | ||
| + | <!-- | ||
==Exercises== | ==Exercises== | ||
<section begin=exercises /> | <section begin=exercises /> | ||
| Line 26: | Line 30: | ||
| + | | ||
==References== | ==References== | ||
<references /> | <references /> | ||
| + | | ||
--> | --> | ||
==Further reading and resources== | ==Further reading and resources== | ||
{{WWW|WWW_ KEGG}} | {{WWW|WWW_ KEGG}} | ||
| − | {{#pmid: | + | {{#pmid:18025687}} |
{{#pmid:19307239}} | {{#pmid:19307239}} | ||
| Line 39: | Line 45: | ||
[[Category:Computational_Systems_Biology]] | [[Category:Computational_Systems_Biology]] | ||
| − | |||
</div> | </div> | ||
Revision as of 21:15, 29 January 2012
KEGG
The Kyoto Encyclopedia of Genes and Genomes (KEGG) is a resource that integrates genomic, chemical and systemic functional information.
Contents
...
Introductory reading
| Kanehisa et al. (2012) KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res 40:D109-14. (pmid: 22080510) |
|
[ PubMed ] [ DOI ] Kyoto Encyclopedia of Genes and Genomes (KEGG, http://www.genome.jp/kegg/ or http://www.kegg.jp/) is a database resource that integrates genomic, chemical and systemic functional information. In particular, gene catalogs from completely sequenced genomes are linked to higher-level systemic functions of the cell, the organism and the ecosystem. Major efforts have been undertaken to manually create a knowledge base for such systemic functions by capturing and organizing experimental knowledge in computable forms; namely, in the forms of KEGG pathway maps, BRITE functional hierarchies and KEGG modules. Continuous efforts have also been made to develop and improve the cross-species annotation procedure for linking genomes to the molecular networks through the KEGG Orthology system. Here we report KEGG Mapper, a collection of tools for KEGG PATHWAY, BRITE and MODULE mapping, enabling integration and interpretation of large-scale data sets. We also report a variant of the KEGG mapping procedure to extend the knowledge base, where different types of data and knowledge, such as disease genes and drug targets, are integrated as part of the KEGG molecular networks. Finally, we describe recent enhancements to the KEGG content, especially the incorporation of disease and drug information used in practice and in society, to support translational bioinformatics. |
Further reading and resources
| KEGG [ link ] [ page ] The Kyoto Encyclopedia of Genes and Genomes (KEGG) is a deeply curated resource that integrates genomic, chemical and systemic functional information. Regrettably, ftp access to this resource is no longer free and personal-use academic licenses outside of Japan are required to purchase a license at the cost of USD 2,000 per year (as per January 2012) from a non-profit organization, founded to ensure the long-term survival of the database. (Read here about the background that explains how public funding is falling short of sustainable levels, thus jeopardizing the ongoing curation and software development activities - and thereby the entire investment into the resource). As of now, use of the Web resources is not to affected. KEGG contains several sections of systems-, genome- and small molecule related information. See here for an overview. The URL links to the pathway map of the yeast cell-cycle. | 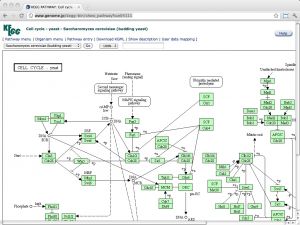 |
| Aoki-Kinoshita & Kanehisa (2007) Gene annotation and pathway mapping in KEGG. Methods Mol Biol 396:71-91. (pmid: 18025687) |
|
[ PubMed ] [ DOI ] KEGG is a database resource (http://www.genome.jp/kegg/) that provides all knowledge about genomes and their relationships to biological systems such as cells and whole organisms as well as their interactions with the environment. KEGG is categorized in terms of building blocks in the genomic space, known as KEGG GENES, the chemical space, KEGG LIGAND, as well as wiring diagrams of interaction and reaction networks, known as KEGG PATHWAY. A fourth database called KEGG BRITE was also recently incorporated to provide computerized annotations and pathway reconstruction based on the current KEGG knowledgebase. KEGG BRITE contains KEGG Orthology (KO), a classification of ortholog and paralog groups based on highly confident sequence similarity scores, and the reaction classification system for biochemical reaction classification, along with other classifications for compounds and drugs. BRITE is also the basis for the KEGG Automatic Annotation Server (KAAS), which automatically annotates a given set of genes and correspondingly generates pathway maps. This chapter introduces KEGG and its various tools for genomic analyses, focusing on the usage of the KEGG GENES, PATHWAY, and BRITE resources and the KAAS tool (see Note 1). |
| Zhang & Wiemann (2009) KEGGgraph: a graph approach to KEGG PATHWAY in R and bioconductor. Bioinformatics 25:1470-1. (pmid: 19307239) |
|
[ PubMed ] [ DOI ] MOTIVATION: KEGG PATHWAY is a service of Kyoto Encyclopedia of Genes and Genomes (KEGG), constructing manually curated pathway maps that represent current knowledge on biological networks in graph models. While valuable graph tools have been implemented in R/Bioconductor, to our knowledge there is currently no software package to parse and analyze KEGG pathways with graph theory. RESULTS: We introduce the software package KEGGgraph in R and Bioconductor, an interface between KEGG pathways and graph models as well as a collection of tools for these graphs. Superior to existing approaches, KEGGgraph captures the pathway topology and allows further analysis or dissection of pathway graphs. We demonstrate the use of the package by the case study of analyzing human pancreatic cancer pathway. AVAILABILITY: KEGGgraph is freely available at the Bioconductor web site (http://www.bioconductor.org). KGML files can be downloaded from KEGG FTP site (ftp://ftp.genome.jp/pub/kegg/xml). |