Difference between revisions of "BIO Assignment 4 2011"
| Line 1: | Line 1: | ||
| − | + | {{Template:Active}} | |
| − | {{Template:Inactive}} | + | <!-- {{Template:Inactive}} --> |
| Line 18: | Line 18: | ||
| | ||
| − | Where is the hidden beauty in structure, and where, the "ultimate truth"? In the previous assignments we have studied sequence conservation in APSES family domains and | + | Where is the hidden beauty in structure, and where, the "ultimate truth"? In the previous assignments we have studied sequence conservation in APSES family domains and we have discovered homologues in all fungal species. This is an ancient protein family that had already duplicated to several paralogues at the time the cenancestor of all fungi lived, more than 600,000,000 years ago, in the [http://www.ucmp.berkeley.edu/fungi/fungifr.html Vendian period] of the Proterozoic era of Precambrian times. |
| − | In order to understand how | + | In order to understand how specific residues in the sequence contribute to the putative function of the protein, and why and how they are conserved throughout evolution, we would need to study an explicit molecular model of an APSES domain protein, bound to its cognate DNA sequence. Explanations of a protein's observed properties and functions can't rely on the general fact that it binds DNA, we need to consider details in terms of specific residues and their spatial arrangement. In particular, it would be interesting to correlate the conservation patterns of key residues with their potential to make specific DNA binding interactions. Unfortunately, no APSES domain structures in complex with bound DNA has been solved up to now, and the experimental evidence we have considered in Assignment 2 ([http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=pubmed&dopt=Abstract&list_uids=10747782 Taylor ''et al.'', 2000]) is not sufficient to unambiguously define the details of how a DNA double helix might be bound. Moreover, at least two distinct modes of DNA binding are known for proteins of the winged-helix superfamily, of which the APSES domain is a member. |
| − | ''In this assignment you will construct a molecular model of the Mbp1 orthologue in your assigned organism, identify similar structures of distantly related domains for which protein-DNA complexes are known, | + | ''In this assignment you will (1) construct a molecular model of the Mbp1 orthologue in your assigned organism, (2) identify similar structures of distantly related domains for which protein-DNA complexes are known, (3) assemble a hypothetical complex structure and(4) discuss whether the available evidence allows you to distinguish between different modes of ligand binding, '' |
For the following, please remember the following terminology: | For the following, please remember the following terminology: | ||
| Line 34: | Line 34: | ||
| | ||
| − | A brief overview article on the construction and use of homology models is linked to the resource section at the bottom of this page. That section also contains links to other sites and resources you might | + | A brief overview article on the construction and use of homology models is linked to the resource section at the bottom of this page. That section also contains links to other sites and resources you might find useful or interesting. |
{{Template:Preparation| | {{Template:Preparation| | ||
| Line 40: | Line 40: | ||
num=4| | num=4| | ||
ord=fourth| | ord=fourth| | ||
| − | due = Monday, November | + | due = Monday, November 12 at 10:00 in the morning}} |
| Line 47: | Line 47: | ||
</div> | </div> | ||
| − | |||
| − | |||
<div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | <div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | ||
| − | === | + | ===(1.1) Template choice and sequence (1 mark)=== |
</div> | </div> | ||
<br> | <br> | ||
| − | Often more than one related structure can be found in the PDB. We have | + | Often more than one related structure can be found in the PDB. We have touched on principles of selecting template structures in the lecture and there is a short summary of [[Template_choice_principles|template choice principles]] on this Wiki. One can either search the PDB itself through its '''Advanced Search''' interface; for example one can search for sequence similarity with a BLAST search, or search for structural similarity by accessing structures according to their CATH or SCOP classification. But one can always also use the BLAST interface at the NCBI, since the sequences contained in PDB files are accessible as a database subsection on the BLAST menu. |
| − | *Use BLAST to identify all PDB files | + | <div style="padding: 5px; background: #DDDDEE;"> |
| + | *Use the NCBI BLAST interface to identify all PDB files that are clearly homologous to your target APSES domain, if you haven't already done so in Assignment 2. Document that you have searched in the correct subsection of the database by selecting "pdb" on the database options menu. For the hits you find, consider how these coordinate sets differ and which features would make each more or less suitable for your task by commenting briefly on | ||
| + | :*sequence similarity to your target | ||
| + | :*size of expected model (length of alignment) | ||
| + | :*presence or absence of ligands | ||
| + | :*experimental method and quality of the data set | ||
| + | Then choose the '''template''' you consider the most suitable and note why you have decided to use this template. | ||
| + | |||
| + | * Retrieve the most suitable template structure coordinate file from the PDB. | ||
| − | + | (0.5 marks) | |
| + | </div> | ||
| − | + | It is not straightforward at all how to number sequence in such a project. The "natural" numbering starts with the start-codon of the full length protein and goes sequentially from there. However, this does not map exactly to other numbering schemes we have encountered. As you know the first residue of the APSES domain as the CDD defines it is not Residue 1 of the Mbp1 protein. The first residue of the e.g. 1MB1 FASTA file '''is''' the first residue of Mbp1 protein, but the last five residues are an artifical His tag. Is H125 of 1MB1 thus equivalent to R125 in MBP1_SACCE? The N-terminus of the Mbp1 crystal structure is disordered. The first residue in the structure is ASN 3, therefore N is the first residue in a FASTA sequence derived from the cordinate section of the PDB file (the <code>ATOM </code> records; whereas the SEQRES records start with MET ... and so on. You need to remember: a sequence number is not absolute, but derived from a particular context. | |
| − | |||
| − | : | ||
| − | + | The homology model will be based on an alignment of target and template. Thus we have to define the target sequence. As discussed in class, PDB files have an explicit and an implied sequence and these do not necessarily have to be the same. To compare the implied and the explicit sequence for the template, you need to extract sequence information from coordinates. One way to do this is via the Web interface for [http://swift.cmbi.ru.nl/servers/html/index.html '''WhatIf'''], a crystallography and molecular modeling package that offers many useful tools for coordinate manipulation tasks. | |
| − | * | + | <div style="padding: 5px; background: #DDDDEE;"> |
| + | *Navigate to the '''Administration''' sub-menu of the [http://swift.cmbi.ru.nl/servers/html/index.html WhatIf Web server]. Follow the link to '''Make sequence file from PDB file'''. Enter the PDB-ID of your template into the form field and '''Send''' the request to the server. The server accesses the PDB file and extracts sequence information directly from the <code>ATOM </code> records of the file. The results will be returned in PIR format. Copy the results, edit them to FASTA format and save them in a text-only file. Make sure you create a valid FASTA formatted file! Use this '''implied''' sequence to check if and how it differs from the sequence ... | ||
| − | + | :*... listed in the <code>SEQRES</code> records of the coordinate file; | |
| + | :*... given in the FASTA sequence for the template, which is provided by the PDB; | ||
| + | :*... stored in the protein database of the NCBI. | ||
| + | : and record your results. | ||
| + | * In a table, establish how the sequence numbers in the coordinate section of your template(*) correspond to your target sequence numbering. | ||
| + | (0.5 marks) | ||
| + | </div> | ||
| + | |||
| + | :(*) <small>These residue numbers are important, since they are referenced e.g. by VMD when you visualize the structure. The easiest way to list them is via the ''Sequence Viewer'' extension of VMD.</small>. | ||
| + | :<small>Don't do this for every residue individually but define ranges. Look at the correspondence of the first and last residue of target and template sequence and take indels into account. Establishing sequence correspondence precisely is crucially important! For example, when a publication refers to a residue by its sequence number, you have to be able to relate that number to the residue numbers of the model as well as your target sequence.</small>. | ||
| | ||
| | ||
<div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | <div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | ||
| − | === The input alignment (1 | + | |
| + | ===(1.2) The input alignment (1 mark)=== | ||
</div> | </div> | ||
<br> | <br> | ||
| − | The sequence alignment between target and template is the single most important factor that determines the quality of your model. | + | The sequence alignment between target and template is the single most important factor that determines the quality of your model. No comparative modeling process will repair an incorrect alignment; it is useful to consider a homology model rather like a three-dimensional map of a sequence alignment rather than a structure in its own right. In a homology modeling project, typically the largest amount of time should be spent on preparing the best possible alignment. Even though automated servers like the SwissModel server will align sequences and select template structures for you, it would be unwise to use these only because they are convenient. You should take advantage of the much more sophisticated alignment methods available. Analysis of wrong models can't be expected to produce right results. |
| − | + | The best possible alignment is usually constructed from a multiple sequence alignment that includes at least '''the target and template sequence''' and other related sequences as well. The additional sequences are an important aid in identifying the correct placement of insertions and deletions. Your alignment should have been carefully reviewed by you and wherever required, manually adjusted to move insertions or deletions between target and template out of the secondary structure elements of the template structure. | |
| − | + | In the case of Mbp1 genes however, all orthologues we have considered have no indels in the APSES domain regions. Evolutionary pressure on the APSES domains has selected against indels in the more than 600 million years these sequences have evolved independently in their respective species. | |
| − | + | Accordingly, all we need to do is to write the APSES domain sequences one under the other. | |
| − | + | <div style="padding: 5px; background: #DDDDEE;"> | |
| − | + | * Copy the FASTA formatted sequence for the APSES domain of your organism's Mbp1 orthologue from the sequences [[All_APSES_domains|defined in Assignment 3]] and save it as FASTA formatted text file. This is your '''target''' sequence. Compare this with the FASTA formatted file you have extracted from the PDB coordinate set. This is your '''template''' sequence. Then generate a multi-FASTA formatted file that contains both sequences, and '''pad''' the sequence(s) where required with hyphens as gap characters, so that target and template sequences have exactly the same length and are aligned. Refer to the [[Assignment_4_fallback_data|'''Fallback data''']] if you are not sure about the format. | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| + | (1 mark) | ||
| + | </div> | ||
| + | <br> | ||
<br> | <br> | ||
| − | + | <div style="padding: 5px; background: #BDC3DC; border:solid 1px #AAAAAA;"> | |
| − | + | ==(2) Homology model== | |
| + | </div> | ||
| + | | ||
| + | | ||
| − | + | <div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | |
| + | === (2.1) SwissModel (1 mark)=== | ||
| + | </div> | ||
| + | <br> | ||
| + | Access the Swissmodel server at '''http://swissmodel.expasy.org''' . Navigate to the '''Alignment Interface'''. | ||
| − | < | + | <br><div style="padding: 5px; background: #DDDDEE;"> |
| − | < | + | *Paste your alignment for target and model into the form field. Refer to the [[Assignment_4_fallback_data|'''Fallback Data file''']] if you are not sure about the format. Make sure to select the correct option for the alignment input format on the form. |
| − | + | :<small>(You have to choose the correct format, and, if e.g. you choose a CLUSTAL format, you have to include a header line and a blank line. In the past we have seen problems with uploading alignments that have not been saved as "text only" and including periods i.e. "." in sequence names of CLUSTAL formatted alignments. Underscores appear to be safe.</small> | |
| − | |||
| − | </ | ||
| − | < | + | * Click '''submit alignment ''' and on the returned page define your '''target''' and '''template''' sequence. For the '''template sequence''' define the PDB ID of the coordinate file. Enter the correct Chain-ID. |
| − | + | :<small>Recently the PDB has undergone a "remediation" process in which archived coordinate files were altered by the database to conform to new format standards. One of the changes was to assign a chain identifier of "A" to all chains that did not previously have a chain identifier. SwissModel uses a derivative of coordinate sets from the PDB (a dataset they call ExPDB). Apparently the PDB proper and ExPDB have now gone out of synchrony; when I entered the (correct, according to PDB) chain designation "A" for 1MB1, SwissModel rejected the alignment with a nondescript error message. When I entered an underscore "_" instead, which would be the designation for a chain without explicit chain identifier, such as the pre-remidation versio of the coordinates, the alignment was accepted and processed. I have e-mailed SwissModel about the problem; they are in the process of correcting it and may or may not be done while you are working on your assignments. If your template chain has the chain identifier "A" and your alignment gets rejected, try entering entering an underscore instead.</small> | |
| − | + | :<small>'''Enter''' the correct chain ID into the form-field even if you think it already appears there, don't simply accept the preloaded default. There is a bug in SwissModel's parser code that may cause incorrect strings to be sent to the server from that field. I have e-mailed SwissModel about the problem which may or may not be corrected while you are working on your assignments.</small> | |
| − | </ | ||
| − | + | *Click '''submit alignment''' and review the alignment on the returned page. Make sure it has been interpreted correctly by the server. The conserved residues have to be lined up and matching. Then click '''submit alignment''' again, to start the modeling process. | |
| − | |||
| − | |||
| − | |||
| − | + | * The resulting page returns information about the resulting model. Save the '''model coordinates''' on your computer. Read the information on what is being returned by the server (click on the red questionmark icon). Paste the Anolea profile into your assignment. | |
| − | + | :<small>Do not paste a screenshot of the result, but copy and paste the image from the Web-page! You do not need to submit the actual coordinate files with your assignment.</small> | |
| − | |||
| − | </ | ||
| − | + | (1 mark) | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
</div> | </div> | ||
<br> | <br> | ||
| + | In case you do not wish to submit the modelling job yourself, or have insurmountable problems when using the SwissModel interface, you may access the result files from the [[Assignment_4_fallback_data|'''Fallback Data file''']]. Document the problems and note this in your assignment. | ||
| − | |||
| − | > | + | <div style="padding: 5px; background: #BDC3DC; border:solid 1px #AAAAAA;"> |
| − | |||
| − | |||
| + | ==(3) Model analysis== | ||
| + | </div> | ||
| | ||
| | ||
| − | <div style="padding: 5px; background: # | + | <div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> |
| + | === (3.1) The PDB file (1 mark)=== | ||
| + | </div> | ||
| + | <br> | ||
| + | |||
| + | Open your '''model''' coordinates in a text-editor (make sure you view the PDB file in a fixed-width font) and consider the following questions: (Alternatively, view the coordinates linked to the [[Assignment_5_fallback_data|'''Fallback Data file''']].) | ||
| − | = | + | <br><div style="padding: 5px; background: #DDDDEE;"> |
| + | *What is the residue number of the first residue in the '''model'''? What should it be, based on the alignment? If the putative DNA binding region was reported to be residues 50-74 in the Mbp1 protein, which residues of the '''model''' correspond to that? | ||
| + | (1 mark) | ||
</div> | </div> | ||
| + | |||
| + | <!-- discuss flagging of loops - setting of B-factor to 99.0 phps. ANOLEA vs. Gromos ... packing vs. energy? --> | ||
| | ||
| | ||
<div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | <div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | ||
| − | === (2 | + | ===(3.2) First visualization (1 mark)=== |
</div> | </div> | ||
<br> | <br> | ||
| − | + | In assignment 2 you have already studied a Mbp1 structure and compared it with your organism's Mbp1 APSES domain, Since a homology model inherits its structural details from the '''template''', the model should look very similar to the original structure but contain the sequence of the '''target'''. | |
| − | <br><div style="padding: 5px; background: # | + | <br><div style="padding: 5px; background: #DDDDEE;"> |
| − | * | + | *Save your '''model''' coordinates to your harddisk and visualize the structure in VMD. (Alternatively, copy and save the coordinates linked to the [[Assignment_4_fallback_data|'''Fallback Data file''']] to your harddisk.) Make an informative stereo view that shows the general orientation of the helix-turn-helix motif and the "wing", and paste it into your assignment. |
| − | |||
| − | * | + | * Discuss briefly which parts of the model may be unreliable and color these (if any) distinctly in your submitted image. |
| − | + | (1 mark) | |
| − | |||
| − | |||
</div> | </div> | ||
<br> | <br> | ||
| − | + | <br> | |
| − | |||
<div style="padding: 5px; background: #BDC3DC; border:solid 1px #AAAAAA;"> | <div style="padding: 5px; background: #BDC3DC; border:solid 1px #AAAAAA;"> | ||
| − | ==( | + | ==(4) The DNA ligand== |
</div> | </div> | ||
| | ||
| Line 276: | Line 193: | ||
<div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | <div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | ||
| − | === ( | + | |
| + | ===(4.1) Finding a similar protein-DNA complex (1 mark)=== | ||
</div> | </div> | ||
<br> | <br> | ||
| − | + | One of the really interesting questions we can discuss with reference to our model is how sequence variation might be converted into changing DNA recognition sites, and then lead to changed cognate DNA binding sequences. But in order to address this, we would need to add a plausible model for how DNA is bound to APSES domains. | |
| − | + | Since there is currently no software available that would accurately model such a complex from first principles, we will base a model of a bound complex on homology modeling as well. This means we need to find a similar structure for which the position of bound DNA is known, then superimpose that structure with our model. This places the DNA molecule into the spatial context of the model we are studying. However, you may remember from the third assignment that the APSES domains in fungi seem to be a relatively small family. And there is no structure available of an APSES domain-DNA complex. How can we find a coordinate set of a strcturally similar protein-DNA complex? | |
| − | + | ||
| − | + | Remember that homologous sequences can have diverged to the point where their sequence similarity is no longer recognizable, however their structure may be quite well conserved. Thus if we could find similar structures in the PDB, these might provide us with some plausible hypotheses for how DNA is bound by APSES domains. We thus need a tool similar to BLAST, but not for the purpose of sequence alignment, but for structure alignment. A kind of BLAST for structures. Just like with sequence searches, we might not want to search with the entire protein, if we are interested in is a subdomain that binds to DNA. Attempting to match all structural elements in addition to the ones we are actually interested in is likely to make the search less specific - we would find false positives that are similar to some irrelevant part of our structure. However, defining too small of a subdomain would also lead to a loss of specificity: in the extreme it is easy to imagine that the search for e.g. a single helix would retrieve very many hits that would be quite meaningless. | |
| + | |||
| + | At the '''NCBI''', [http://www.ncbi.nlm.nih.gov/Structure/VAST/vast.shtml VAST] is provided as a search tool for structural similarity search. | ||
| − | + | At the '''EBI''' there are a number of very well designed structure analysis tools linked off the [http://www.ebi.ac.uk/Tools/structural.html '''Structural Analysis''' page]. As part of its MSD Services, [http://www.ebi.ac.uk/msd-srv/ssm/ '''MSDfold'''] provides a convenient interface for structure searches. | |
| − | + | However we have also read previously that the APSES domains are members of a much larger superfamily, the "winged helix" DNA binding domains , of which hundreds of structures have been solved. | |
| − | |||
| − | |||
| − | |||
| − | |||
<br> | <br> | ||
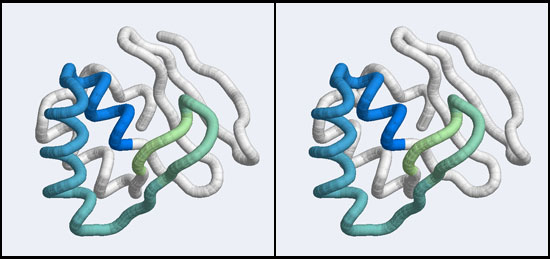
| − | + | [[Image:A5_Mbp1_subdomain.jpg|frame|none|Stereo-view of a subdomain within the 1MB1 structure that includes residues 36 to 76. The color gradient ramps from blue (36) to green (76) and the "wing" is clearly seen as the green pair of beta-strands, extending to the right of the helix-turn-helix motif.]] | |
| − | <br> | + | <br> |
| − | |||
| − | + | APSES domains represent one branch of the tree of helix-turn-helix (HTH) DNA binding modules. (A recent review on HTH proteins is linked from the resources section at the bottom of this page). Winged Helix domains typically bind their cognate DNA with a "recognition helix" which precedes the beta hairpin and binds into the major groove; additional stabilizing interactions are provided by the edge of a beta-strand binding into the minor groove. This is good news: once we have determined that the APSES domain is actually an example of a larger group of transcription factors, we can compare our model to a structure of a protein-DNA complex. CATH does not provide information on complexes, but we can search the PDB with CATH codes in the following way: | |
| − | |||
| + | * Access [http://cathwww.biochem.ucl.ac.uk/cgi-bin/cath/GotoCath.pl?cath=1.10.10.10 CATH domain 1.10.10.10]. | ||
| + | * Navigate to the [http://www.pdb.org/ PDB home page] and follow the link to [http://www.pdb.org/pdb/search/advSearch.do Advanced Search] | ||
| + | * In the options menu for "Choose a Query Type" select Structure Features → CATH classification. A window will open that allows you to navigate down through the CATH tree. The interface is awkward because it does not display the actual CATH codes along with the class names, but you can view the class names on the CATH page linked above. Click on '''the triangle icons''' before "Mainly Alpha"→"Orthogonal Bundle"→"ARC repressor mutant, subunit A" then click on the link to "winged helix repressor DNA binding domain". As of this writing, this subquery matches 295 structures. | ||
| + | * Click on the (+) button behind the subquery to add an additional query. Select the option "Structure Summary"→"Molecule / Chain type". In the option menus that pop up, select "Contains Protein → Yes", "Contains DNA → Yes""Contains RNA → Ignore". This selects files that contain Protein-DNA complexes. | ||
| + | * Check the box below this subquery to "Remove Similar Sequences at 90% identity" and click on "Evaluate Query". As of this writing, seventy complexes were returned. | ||
| + | * In the left-hand menu, under the Tabulate section, click on the "Collage" function to display icons of the structure files. This is a fast way to obtain an overview of the structures that have been returned. First of all you may notice that in fact not all of the structures are really different, despite selecting only to retrieve dissimilar sequences. This appears to be a deficiency of the algorithm. But you can also easily recognize how the recognition helix inserts into the major groove of most of the structures that were returned (at least those where the domain is not a very small part of a much larger complex). There is one exception: the structure 1DP7 shows how the human RFX1 protein binds DNA in a non-canonical way. We shall use structural superposition of your homology model and two of the winged-helix proteins to decide which mode of DNA binding seems to be more plausible for Mbp1 homologues. | ||
| − | + | <br><div style="padding: 5px; background: #DDDDEE;"> | |
| + | * Follow the procedure outlined above, from a CATH entry page up to viewing a Collage (or alternatively a tabular view) of the retrieved coordinate files. You can be maximally concise documenting the procedure I have defined above, but do spend a bit of time to understand the key elements of the PDB's advanced search interface. | ||
| − | + | (1 mark) | |
| − | + | </div> | |
<div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | <div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | ||
| − | ===( | + | ===(4.2) Preparation and superposition of a canonical complex (1 mark)=== |
</div> | </div> | ||
<br> | <br> | ||
| − | The | + | The structure we shall use as a reference for the canonical binding mode is the Elk-1 transcription factor. |
| − | + | [[Image:A5_canonical_wHTH.jpg|frame|none|Stereo-view of the canonical DNA binding mode of the Winged Helix domain family. Shown here is the Elk-1 transcription factor - an ETS DNA binding domain - in complex with a high-affinity binding site (1DUX). Note how the "recognition helix" inserts into the major groove of the DNA molecule. The color gradient ramps from blue (34) to green (84). Note how the first helix of the "helix-turn-helix" architecture serves only to position the recognition helix and makes few interactions by itself.]] | |
| − | + | The 1DUX coordinate-file contains two protein domains and two B-DNA dimers in one asymmetric unit. For simplicity, let's delete the second copy. | |
| − | + | * Access the PDB and navigate to the 1DUX structure explorer page. Download the coordinates to your computer. | |
| + | * Open the coordinate file in a text-editor and delete the coordinates for chains <code>D</code>,<code>E</code> and <code>F</code>; you may also delete all <code>HETATM</code> records and the <code>MASTER</code> record. Save the file with a different name, e.g. 1DUX_monomer.pdb . | ||
| + | * Open VMD and load your homology model. Turn off the axes, display the model as a Tube representation in stereo, and color it by Index. Then load your edited 1DUX file, display this coordinate set in a tube representation as well, and color it by ColorID in some color you like. It is important that you can distinguish easily which structure is which | ||
| + | * You could use the Extensions→Analysis→RMSD calculator interface to superimpose the two strutcures '''IF''' you would know which residues correspond to each other. Sometimes it is useful to do exactly that: define exact correspondences between residue pairs and superimpose according to these selected pairs. For our purpose it is much simpler to use the Multiseq tool (and the structures are simple and small enough that the STAMP algorithm for structural alignment can define corresponding residue pairs automatically). Open the '''multiseq''' extension window, select the check-boxes next to both protein structures, and open the '''Tools→Stamp Structural Alignment''' interface. | ||
| + | * In the "'Stamp Alignment Options'" window, check the radio-button for ''Align the following ...'' '''Marked Structures''' and click on '''OK'''. | ||
| + | * In the '''Graphical Representations''' window, double-click on all "NewCartoon" representations for both molecules, to undisplay them. | ||
| + | * You should now see a superimposed tube model of your homology model and the 1DUX protein-DNA complex. You can explore it, display side-chains etc. and study some of the details of how a transcription factor recognizes and binds to its cognate DNA sequence. However, remember that the model's side-chain orientations have not been experimentally determined but inferred from the template, and that the template's strcture was determined in the absence of bound ligand. | ||
| − | + | <br><div style="padding: 5px; background: #DDDDEE;"> | |
| + | * Orient and scale your superimposed structures so that their structural similarity is apparent, and the recognition helix can be clearly seen inserting into the DNA major groove. Paste a copy of your image into your assignment. Remark briefly on which parts of the structure appear to superimpose best. Note whether this orientation of a B-DNA double-helix is a plausible model for DNA binding of your Mbp1 orthologue. | ||
| − | + | (1 mark) | |
| + | </div> | ||
| + | <br> | ||
| + | | ||
| − | |||
| − | < | + | <div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> |
| − | + | ===(4.2) Preparation and superposition of a non-canonical complex (1 mark)=== | |
| + | </div> | ||
| + | <br> | ||
| − | + | The structure displaying a non-canonical complex between a winged-helix domain and its cognate DNA binding site is the human Regulatory Factor X. | |
| − | [[Image: | + | [[Image:A5_non-canonical_wHTH.jpg|frame|none|Stereo-view of a non-canonical wHTH-DNA complex, discovered in with the stucture of human Regulatory Factor X (hRFX) binding its cognate X-box DNA sequence (1DP7). Note how the helix that coressponds to the recogition helix in the canonical domain lies across the minor groove whereas the beta-"wing" inserts into the major groove. The color gradient ramps from blue (18) to green (68).]] |
| − | + | The 1DP7 coordinate-file contains only one protein domain and only one B-DNA monomer in its asymmetric unit. This is a file for which we have to generate ''biological unit'' coordinates! Then, for simplicity we will delete the second protein monomer. As you know, there are at least two systems that make the so-called biological units available: the PDB itself, through the Biological Unit file that is accessible via the "Download Files" section on any Structure Explorer page, and the EBI through the PQS service. '''How''' the biological units are stored is subtly different for both cases and for our purpose this small difference is important. The PDB generates additional chins as copies of the original and delineates them with <code>MODEL</code>, <code>ENDMDL</code> records, just like in a multi-structure NMR file. The chain IDs and the atom numbers are the same as the original. The EBI's PQS service creates copies that have distinct atomnumbers and chain IDs. The difference is that the PDB file thus '''contains the same molecule in two different orientations''', wheras the PQS file contains '''two independent molecules'''. This is an important difference when it comes to selecting residues, visualizing and superimposing structures. For VMD, the PQS way of doing things is the right way to go, since by default only the first <code>MODEL</code> will be displayed if several are available. | |
| − | + | * Access the [http://pqs.ebi.ac.uk/ '''EBI PQS server'''], enter 1DP7 into the '''PDBidcode''' form field and click on '''Submit'''. | |
| − | + | * On the results page, click on the link under '''1dp7_0''', which is the unique suggestion for a biological unit that the server has identified. | |
| + | * On the PQS OUTPUT page that is retrieved, click on the '''1dp7.mmol''' link, this will load the PDB formatted coordinate file. | ||
| + | * Save the coordinates as 1DP7_complex.pdb (or some other name that makes sense to you), open it in a text editor, delete the <code>HETATM</code> records from the end and the entire chain "B". Also make sure not to delete any of the <code>TER</code> records for chains "D", "P" or "A". Save the file. | ||
| + | * In the multiseq window, choose File→Import Data, '''Browse...''' to your 1DP7_complex file, select it and click on '''Open'''. Click '''OK''' to load the file. | ||
| + | * Mark all three protein chains by selecting the checkbox next to thier name and again run the STAMP structural alignment. | ||
| + | * In the graphical representations window, double-click again on all cartoon representations that multiseq has generated to undisplay them, undisplay also the Tube representation of 1DUX, create a Tube representatrion for 1DP7, and select a Color by ColorID (a differnet color you like). The resulting scene should look similar to the one you have created above, only with 1DP7 in place of 1DUX and colored differently. | ||
| − | * | + | <br><div style="padding: 5px; background: #DDDDEE;"> |
| − | + | * Orient and scale your superimposed structures so that their structural similarity is apparent, the orientation is similar to the scene generated above and the 1DP7 "wing" can be clearly seen inserting into the DNA major groove. Paste a copy of your image into your assignment. Remark briefly on which parts of the structure appear to superimpose best. Note whether this orientation of a B-DNA double-helix is a plausible model for DNA binding of your Mbp1 orthologue. | |
| − | |||
| − | |||
| − | |||
| − | + | (1 mark) | |
| + | </div> | ||
| + | <br> | ||
| + | <div style="padding: 5px; background: #E9EBF3; border:solid 1px #AAAAAA;"> | ||
| − | + | ===(4.3) Interpretation (2 marks)=== | |
| − | |||
</div> | </div> | ||
<br> | <br> | ||
| − | |||
| + | In your previous assignment, you have commented on conservation patterns in Mbp1 orthologues. You can refer back to your last results (easier to do), or you can import the APSES domain alignment for Mbp1 proteins and again color by conservation (easier to study) to briefly discuss the following question. | ||
| + | |||
| + | <br><div style="padding: 5px; background: #DDDDEE;"> | ||
| + | * Considering the conservation patterns for Mbp1 orthologues, and assuming that all these orthologues bind DNA in a similar way, which model appears to be more plausible for protein-DNA interactions in APSES domains? Is it the canonical, or the non-canonical binding mode? Discuss briefly what you would expect to find and how this relates to your observations. Distinguish clearly between experimental evidence, computational inference and empirical hypothesis. You are of course welcome to paste detail views (stereo !) of particular sidechains, or surfaces etc. if this helps your arguments. Sometimes a picture is worth many words. But this is not a requirement, we are more interested in evidence-based reasoning than in the form of the presentation. | ||
| + | |||
| + | (2 marks) | ||
| + | </div> | ||
| + | <br> | ||
| + | <br> | ||
<div style="padding: 5px; background: #BDC3DC; border:solid 1px #AAAAAA;"> | <div style="padding: 5px; background: #BDC3DC; border:solid 1px #AAAAAA;"> | ||
| − | ==( | + | ==(5) Summary of Resources== |
</div> | </div> | ||
<br> | <br> | ||
| − | ;Links | + | ;Links and background reading |
| + | |||
:* [http://biochemistry.utoronto.ca/undergraduates/courses/BCH441H/restricted/Peitsch_2002_UseOfModels.pdf '''Review (PDF, restricted)''' Manuel Peitsch on Homology Modeling] | :* [http://biochemistry.utoronto.ca/undergraduates/courses/BCH441H/restricted/Peitsch_2002_UseOfModels.pdf '''Review (PDF, restricted)''' Manuel Peitsch on Homology Modeling] | ||
| − | :* [http://biochemistry.utoronto.ca/undergraduates/courses/BCH441H/restricted/Aravind_2005_HTHdomains.pdf '''Review (PDF, restricted)''' Aravind ''et al.'' Helix-turn-helix domains] ( | + | :* [http://biochemistry.utoronto.ca/undergraduates/courses/BCH441H/restricted/Aravind_2005_HTHdomains.pdf '''Review (PDF, restricted)''' Aravind ''et al.'' Helix-turn-helix domains] |
| − | :* [[ | + | :* [http://biochemistry.utoronto.ca/undergraduates/courses/BCH441H/restricted/2000_Gajiwala_WingedHelixDomains.pdf '''Review (PDF, restricted)''' Gajiwala & Burley, winged-Helix domains] |
| − | :* [http://www. | + | :* [[Organism_list_2007|Assigned Organisms]] |
| + | :* [http://www.wwpdb.org/documentation/format23/v2.3.html '''PDB file format'''] (see the Coordinate Section if you are unsure about chain identifiers) | ||
:* [http://en.wikipedia.org/wiki/Structural_alignment Wikipedia on '''Structural Superposition'''] <small>(although the article is called "Structural Alignment")</small> | :* [http://en.wikipedia.org/wiki/Structural_alignment Wikipedia on '''Structural Superposition'''] <small>(although the article is called "Structural Alignment")</small> | ||
| − | + | ;[[Assignment_4_fallback_data|'''Fallback Data page''']] | |
;Alignments | ;Alignments | ||
| − | :* [[ | + | :* [[APSES_domains_MUSCLE|APSES domains MUSCLE aligned]] |
| | ||
| | ||
| − | + | {{Template:Assignment_Footer}} | |
| − | |||
| − | |||
| − | |||
| − | |||
Revision as of 13:52, 1 November 2007
Note! This assignment is currently active. All significant changes will be announced on the mailing list.
Contents
Assignment 4 - Homology modeling
- How could the search for ultimate truth have revealed so hideous and visceral-looking an object?
-
- Max Perutz (on his first glimpse of the Hemoglobin structure)
Where is the hidden beauty in structure, and where, the "ultimate truth"? In the previous assignments we have studied sequence conservation in APSES family domains and we have discovered homologues in all fungal species. This is an ancient protein family that had already duplicated to several paralogues at the time the cenancestor of all fungi lived, more than 600,000,000 years ago, in the Vendian period of the Proterozoic era of Precambrian times.
In order to understand how specific residues in the sequence contribute to the putative function of the protein, and why and how they are conserved throughout evolution, we would need to study an explicit molecular model of an APSES domain protein, bound to its cognate DNA sequence. Explanations of a protein's observed properties and functions can't rely on the general fact that it binds DNA, we need to consider details in terms of specific residues and their spatial arrangement. In particular, it would be interesting to correlate the conservation patterns of key residues with their potential to make specific DNA binding interactions. Unfortunately, no APSES domain structures in complex with bound DNA has been solved up to now, and the experimental evidence we have considered in Assignment 2 (Taylor et al., 2000) is not sufficient to unambiguously define the details of how a DNA double helix might be bound. Moreover, at least two distinct modes of DNA binding are known for proteins of the winged-helix superfamily, of which the APSES domain is a member.
In this assignment you will (1) construct a molecular model of the Mbp1 orthologue in your assigned organism, (2) identify similar structures of distantly related domains for which protein-DNA complexes are known, (3) assemble a hypothetical complex structure and(4) discuss whether the available evidence allows you to distinguish between different modes of ligand binding,
For the following, please remember the following terminology:
- Target
- The protein that you are planning to model.
- Template
- The protein whose structure you are using as a guide to build the model.
- Model
- The structure that results from the modeling process. It has the Target sequence and is similar to the Template structure.
A brief overview article on the construction and use of homology models is linked to the resource section at the bottom of this page. That section also contains links to other sites and resources you might find useful or interesting.
Preparation, submission and due date
- Read carefully.
- Be sure you have understood all parts of the assignment and cover all questions in your answers! Sadly, we see too many assignments which, arduously effected, nevertheless intimate nescience of elementary tenets of molecular biology. If the sentence above did not trigger an urge to open a dictionary, you are trying to guess, rather than confirm possibly important information.
Review the guidelines for preparation and submission of BCH441 assignments.
The due date for the assignment is Monday, November 12 at 10:00 in the morning.
(1) Preparation
(1.1) Template choice and sequence (1 mark)
Often more than one related structure can be found in the PDB. We have touched on principles of selecting template structures in the lecture and there is a short summary of template choice principles on this Wiki. One can either search the PDB itself through its Advanced Search interface; for example one can search for sequence similarity with a BLAST search, or search for structural similarity by accessing structures according to their CATH or SCOP classification. But one can always also use the BLAST interface at the NCBI, since the sequences contained in PDB files are accessible as a database subsection on the BLAST menu.
- Use the NCBI BLAST interface to identify all PDB files that are clearly homologous to your target APSES domain, if you haven't already done so in Assignment 2. Document that you have searched in the correct subsection of the database by selecting "pdb" on the database options menu. For the hits you find, consider how these coordinate sets differ and which features would make each more or less suitable for your task by commenting briefly on
- sequence similarity to your target
- size of expected model (length of alignment)
- presence or absence of ligands
- experimental method and quality of the data set
Then choose the template you consider the most suitable and note why you have decided to use this template.
- Retrieve the most suitable template structure coordinate file from the PDB.
(0.5 marks)
It is not straightforward at all how to number sequence in such a project. The "natural" numbering starts with the start-codon of the full length protein and goes sequentially from there. However, this does not map exactly to other numbering schemes we have encountered. As you know the first residue of the APSES domain as the CDD defines it is not Residue 1 of the Mbp1 protein. The first residue of the e.g. 1MB1 FASTA file is the first residue of Mbp1 protein, but the last five residues are an artifical His tag. Is H125 of 1MB1 thus equivalent to R125 in MBP1_SACCE? The N-terminus of the Mbp1 crystal structure is disordered. The first residue in the structure is ASN 3, therefore N is the first residue in a FASTA sequence derived from the cordinate section of the PDB file (the ATOM records; whereas the SEQRES records start with MET ... and so on. You need to remember: a sequence number is not absolute, but derived from a particular context.
The homology model will be based on an alignment of target and template. Thus we have to define the target sequence. As discussed in class, PDB files have an explicit and an implied sequence and these do not necessarily have to be the same. To compare the implied and the explicit sequence for the template, you need to extract sequence information from coordinates. One way to do this is via the Web interface for WhatIf, a crystallography and molecular modeling package that offers many useful tools for coordinate manipulation tasks.
- Navigate to the Administration sub-menu of the WhatIf Web server. Follow the link to Make sequence file from PDB file. Enter the PDB-ID of your template into the form field and Send the request to the server. The server accesses the PDB file and extracts sequence information directly from the
ATOMrecords of the file. The results will be returned in PIR format. Copy the results, edit them to FASTA format and save them in a text-only file. Make sure you create a valid FASTA formatted file! Use this implied sequence to check if and how it differs from the sequence ...
- ... listed in the
SEQRESrecords of the coordinate file; - ... given in the FASTA sequence for the template, which is provided by the PDB;
- ... stored in the protein database of the NCBI.
- ... listed in the
- and record your results.
- In a table, establish how the sequence numbers in the coordinate section of your template(*) correspond to your target sequence numbering.
(0.5 marks)
- (*) These residue numbers are important, since they are referenced e.g. by VMD when you visualize the structure. The easiest way to list them is via the Sequence Viewer extension of VMD..
- Don't do this for every residue individually but define ranges. Look at the correspondence of the first and last residue of target and template sequence and take indels into account. Establishing sequence correspondence precisely is crucially important! For example, when a publication refers to a residue by its sequence number, you have to be able to relate that number to the residue numbers of the model as well as your target sequence..
(1.2) The input alignment (1 mark)
The sequence alignment between target and template is the single most important factor that determines the quality of your model. No comparative modeling process will repair an incorrect alignment; it is useful to consider a homology model rather like a three-dimensional map of a sequence alignment rather than a structure in its own right. In a homology modeling project, typically the largest amount of time should be spent on preparing the best possible alignment. Even though automated servers like the SwissModel server will align sequences and select template structures for you, it would be unwise to use these only because they are convenient. You should take advantage of the much more sophisticated alignment methods available. Analysis of wrong models can't be expected to produce right results.
The best possible alignment is usually constructed from a multiple sequence alignment that includes at least the target and template sequence and other related sequences as well. The additional sequences are an important aid in identifying the correct placement of insertions and deletions. Your alignment should have been carefully reviewed by you and wherever required, manually adjusted to move insertions or deletions between target and template out of the secondary structure elements of the template structure.
In the case of Mbp1 genes however, all orthologues we have considered have no indels in the APSES domain regions. Evolutionary pressure on the APSES domains has selected against indels in the more than 600 million years these sequences have evolved independently in their respective species.
Accordingly, all we need to do is to write the APSES domain sequences one under the other.
- Copy the FASTA formatted sequence for the APSES domain of your organism's Mbp1 orthologue from the sequences defined in Assignment 3 and save it as FASTA formatted text file. This is your target sequence. Compare this with the FASTA formatted file you have extracted from the PDB coordinate set. This is your template sequence. Then generate a multi-FASTA formatted file that contains both sequences, and pad the sequence(s) where required with hyphens as gap characters, so that target and template sequences have exactly the same length and are aligned. Refer to the Fallback data if you are not sure about the format.
(1 mark)
(2) Homology model
(2.1) SwissModel (1 mark)
Access the Swissmodel server at http://swissmodel.expasy.org . Navigate to the Alignment Interface.
- Paste your alignment for target and model into the form field. Refer to the Fallback Data file if you are not sure about the format. Make sure to select the correct option for the alignment input format on the form.
- (You have to choose the correct format, and, if e.g. you choose a CLUSTAL format, you have to include a header line and a blank line. In the past we have seen problems with uploading alignments that have not been saved as "text only" and including periods i.e. "." in sequence names of CLUSTAL formatted alignments. Underscores appear to be safe.
- Click submit alignment and on the returned page define your target and template sequence. For the template sequence define the PDB ID of the coordinate file. Enter the correct Chain-ID.
- Recently the PDB has undergone a "remediation" process in which archived coordinate files were altered by the database to conform to new format standards. One of the changes was to assign a chain identifier of "A" to all chains that did not previously have a chain identifier. SwissModel uses a derivative of coordinate sets from the PDB (a dataset they call ExPDB). Apparently the PDB proper and ExPDB have now gone out of synchrony; when I entered the (correct, according to PDB) chain designation "A" for 1MB1, SwissModel rejected the alignment with a nondescript error message. When I entered an underscore "_" instead, which would be the designation for a chain without explicit chain identifier, such as the pre-remidation versio of the coordinates, the alignment was accepted and processed. I have e-mailed SwissModel about the problem; they are in the process of correcting it and may or may not be done while you are working on your assignments. If your template chain has the chain identifier "A" and your alignment gets rejected, try entering entering an underscore instead.
- Enter the correct chain ID into the form-field even if you think it already appears there, don't simply accept the preloaded default. There is a bug in SwissModel's parser code that may cause incorrect strings to be sent to the server from that field. I have e-mailed SwissModel about the problem which may or may not be corrected while you are working on your assignments.
- Click submit alignment and review the alignment on the returned page. Make sure it has been interpreted correctly by the server. The conserved residues have to be lined up and matching. Then click submit alignment again, to start the modeling process.
- The resulting page returns information about the resulting model. Save the model coordinates on your computer. Read the information on what is being returned by the server (click on the red questionmark icon). Paste the Anolea profile into your assignment.
- Do not paste a screenshot of the result, but copy and paste the image from the Web-page! You do not need to submit the actual coordinate files with your assignment.
(1 mark)
In case you do not wish to submit the modelling job yourself, or have insurmountable problems when using the SwissModel interface, you may access the result files from the Fallback Data file. Document the problems and note this in your assignment.
(3) Model analysis
(3.1) The PDB file (1 mark)
Open your model coordinates in a text-editor (make sure you view the PDB file in a fixed-width font) and consider the following questions: (Alternatively, view the coordinates linked to the Fallback Data file.)
- What is the residue number of the first residue in the model? What should it be, based on the alignment? If the putative DNA binding region was reported to be residues 50-74 in the Mbp1 protein, which residues of the model correspond to that?
(1 mark)
(3.2) First visualization (1 mark)
In assignment 2 you have already studied a Mbp1 structure and compared it with your organism's Mbp1 APSES domain, Since a homology model inherits its structural details from the template, the model should look very similar to the original structure but contain the sequence of the target.
- Save your model coordinates to your harddisk and visualize the structure in VMD. (Alternatively, copy and save the coordinates linked to the Fallback Data file to your harddisk.) Make an informative stereo view that shows the general orientation of the helix-turn-helix motif and the "wing", and paste it into your assignment.
- Discuss briefly which parts of the model may be unreliable and color these (if any) distinctly in your submitted image.
(1 mark)
(4) The DNA ligand
(4.1) Finding a similar protein-DNA complex (1 mark)
One of the really interesting questions we can discuss with reference to our model is how sequence variation might be converted into changing DNA recognition sites, and then lead to changed cognate DNA binding sequences. But in order to address this, we would need to add a plausible model for how DNA is bound to APSES domains.
Since there is currently no software available that would accurately model such a complex from first principles, we will base a model of a bound complex on homology modeling as well. This means we need to find a similar structure for which the position of bound DNA is known, then superimpose that structure with our model. This places the DNA molecule into the spatial context of the model we are studying. However, you may remember from the third assignment that the APSES domains in fungi seem to be a relatively small family. And there is no structure available of an APSES domain-DNA complex. How can we find a coordinate set of a strcturally similar protein-DNA complex?
Remember that homologous sequences can have diverged to the point where their sequence similarity is no longer recognizable, however their structure may be quite well conserved. Thus if we could find similar structures in the PDB, these might provide us with some plausible hypotheses for how DNA is bound by APSES domains. We thus need a tool similar to BLAST, but not for the purpose of sequence alignment, but for structure alignment. A kind of BLAST for structures. Just like with sequence searches, we might not want to search with the entire protein, if we are interested in is a subdomain that binds to DNA. Attempting to match all structural elements in addition to the ones we are actually interested in is likely to make the search less specific - we would find false positives that are similar to some irrelevant part of our structure. However, defining too small of a subdomain would also lead to a loss of specificity: in the extreme it is easy to imagine that the search for e.g. a single helix would retrieve very many hits that would be quite meaningless.
At the NCBI, VAST is provided as a search tool for structural similarity search.
At the EBI there are a number of very well designed structure analysis tools linked off the Structural Analysis page. As part of its MSD Services, MSDfold provides a convenient interface for structure searches.
However we have also read previously that the APSES domains are members of a much larger superfamily, the "winged helix" DNA binding domains , of which hundreds of structures have been solved.
APSES domains represent one branch of the tree of helix-turn-helix (HTH) DNA binding modules. (A recent review on HTH proteins is linked from the resources section at the bottom of this page). Winged Helix domains typically bind their cognate DNA with a "recognition helix" which precedes the beta hairpin and binds into the major groove; additional stabilizing interactions are provided by the edge of a beta-strand binding into the minor groove. This is good news: once we have determined that the APSES domain is actually an example of a larger group of transcription factors, we can compare our model to a structure of a protein-DNA complex. CATH does not provide information on complexes, but we can search the PDB with CATH codes in the following way:
- Access CATH domain 1.10.10.10.
- Navigate to the PDB home page and follow the link to Advanced Search
- In the options menu for "Choose a Query Type" select Structure Features → CATH classification. A window will open that allows you to navigate down through the CATH tree. The interface is awkward because it does not display the actual CATH codes along with the class names, but you can view the class names on the CATH page linked above. Click on the triangle icons before "Mainly Alpha"→"Orthogonal Bundle"→"ARC repressor mutant, subunit A" then click on the link to "winged helix repressor DNA binding domain". As of this writing, this subquery matches 295 structures.
- Click on the (+) button behind the subquery to add an additional query. Select the option "Structure Summary"→"Molecule / Chain type". In the option menus that pop up, select "Contains Protein → Yes", "Contains DNA → Yes""Contains RNA → Ignore". This selects files that contain Protein-DNA complexes.
- Check the box below this subquery to "Remove Similar Sequences at 90% identity" and click on "Evaluate Query". As of this writing, seventy complexes were returned.
- In the left-hand menu, under the Tabulate section, click on the "Collage" function to display icons of the structure files. This is a fast way to obtain an overview of the structures that have been returned. First of all you may notice that in fact not all of the structures are really different, despite selecting only to retrieve dissimilar sequences. This appears to be a deficiency of the algorithm. But you can also easily recognize how the recognition helix inserts into the major groove of most of the structures that were returned (at least those where the domain is not a very small part of a much larger complex). There is one exception: the structure 1DP7 shows how the human RFX1 protein binds DNA in a non-canonical way. We shall use structural superposition of your homology model and two of the winged-helix proteins to decide which mode of DNA binding seems to be more plausible for Mbp1 homologues.
- Follow the procedure outlined above, from a CATH entry page up to viewing a Collage (or alternatively a tabular view) of the retrieved coordinate files. You can be maximally concise documenting the procedure I have defined above, but do spend a bit of time to understand the key elements of the PDB's advanced search interface.
(1 mark)
(4.2) Preparation and superposition of a canonical complex (1 mark)
The structure we shall use as a reference for the canonical binding mode is the Elk-1 transcription factor.
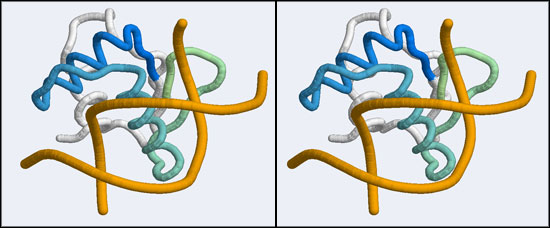
The 1DUX coordinate-file contains two protein domains and two B-DNA dimers in one asymmetric unit. For simplicity, let's delete the second copy.
- Access the PDB and navigate to the 1DUX structure explorer page. Download the coordinates to your computer.
- Open the coordinate file in a text-editor and delete the coordinates for chains
D,EandF; you may also delete allHETATMrecords and theMASTERrecord. Save the file with a different name, e.g. 1DUX_monomer.pdb . - Open VMD and load your homology model. Turn off the axes, display the model as a Tube representation in stereo, and color it by Index. Then load your edited 1DUX file, display this coordinate set in a tube representation as well, and color it by ColorID in some color you like. It is important that you can distinguish easily which structure is which
- You could use the Extensions→Analysis→RMSD calculator interface to superimpose the two strutcures IF you would know which residues correspond to each other. Sometimes it is useful to do exactly that: define exact correspondences between residue pairs and superimpose according to these selected pairs. For our purpose it is much simpler to use the Multiseq tool (and the structures are simple and small enough that the STAMP algorithm for structural alignment can define corresponding residue pairs automatically). Open the multiseq extension window, select the check-boxes next to both protein structures, and open the Tools→Stamp Structural Alignment interface.
- In the "'Stamp Alignment Options'" window, check the radio-button for Align the following ... Marked Structures and click on OK.
- In the Graphical Representations window, double-click on all "NewCartoon" representations for both molecules, to undisplay them.
- You should now see a superimposed tube model of your homology model and the 1DUX protein-DNA complex. You can explore it, display side-chains etc. and study some of the details of how a transcription factor recognizes and binds to its cognate DNA sequence. However, remember that the model's side-chain orientations have not been experimentally determined but inferred from the template, and that the template's strcture was determined in the absence of bound ligand.
- Orient and scale your superimposed structures so that their structural similarity is apparent, and the recognition helix can be clearly seen inserting into the DNA major groove. Paste a copy of your image into your assignment. Remark briefly on which parts of the structure appear to superimpose best. Note whether this orientation of a B-DNA double-helix is a plausible model for DNA binding of your Mbp1 orthologue.
(1 mark)
(4.2) Preparation and superposition of a non-canonical complex (1 mark)
The structure displaying a non-canonical complex between a winged-helix domain and its cognate DNA binding site is the human Regulatory Factor X.
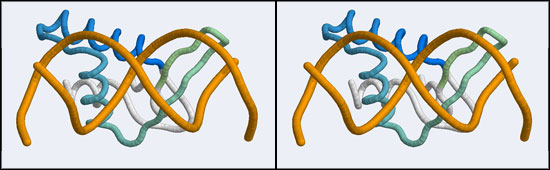
The 1DP7 coordinate-file contains only one protein domain and only one B-DNA monomer in its asymmetric unit. This is a file for which we have to generate biological unit coordinates! Then, for simplicity we will delete the second protein monomer. As you know, there are at least two systems that make the so-called biological units available: the PDB itself, through the Biological Unit file that is accessible via the "Download Files" section on any Structure Explorer page, and the EBI through the PQS service. How the biological units are stored is subtly different for both cases and for our purpose this small difference is important. The PDB generates additional chins as copies of the original and delineates them with MODEL, ENDMDL records, just like in a multi-structure NMR file. The chain IDs and the atom numbers are the same as the original. The EBI's PQS service creates copies that have distinct atomnumbers and chain IDs. The difference is that the PDB file thus contains the same molecule in two different orientations, wheras the PQS file contains two independent molecules. This is an important difference when it comes to selecting residues, visualizing and superimposing structures. For VMD, the PQS way of doing things is the right way to go, since by default only the first MODEL will be displayed if several are available.
- Access the EBI PQS server, enter 1DP7 into the PDBidcode form field and click on Submit.
- On the results page, click on the link under 1dp7_0, which is the unique suggestion for a biological unit that the server has identified.
- On the PQS OUTPUT page that is retrieved, click on the 1dp7.mmol link, this will load the PDB formatted coordinate file.
- Save the coordinates as 1DP7_complex.pdb (or some other name that makes sense to you), open it in a text editor, delete the
HETATMrecords from the end and the entire chain "B". Also make sure not to delete any of theTERrecords for chains "D", "P" or "A". Save the file. - In the multiseq window, choose File→Import Data, Browse... to your 1DP7_complex file, select it and click on Open. Click OK to load the file.
- Mark all three protein chains by selecting the checkbox next to thier name and again run the STAMP structural alignment.
- In the graphical representations window, double-click again on all cartoon representations that multiseq has generated to undisplay them, undisplay also the Tube representation of 1DUX, create a Tube representatrion for 1DP7, and select a Color by ColorID (a differnet color you like). The resulting scene should look similar to the one you have created above, only with 1DP7 in place of 1DUX and colored differently.
- Orient and scale your superimposed structures so that their structural similarity is apparent, the orientation is similar to the scene generated above and the 1DP7 "wing" can be clearly seen inserting into the DNA major groove. Paste a copy of your image into your assignment. Remark briefly on which parts of the structure appear to superimpose best. Note whether this orientation of a B-DNA double-helix is a plausible model for DNA binding of your Mbp1 orthologue.
(1 mark)
(4.3) Interpretation (2 marks)
In your previous assignment, you have commented on conservation patterns in Mbp1 orthologues. You can refer back to your last results (easier to do), or you can import the APSES domain alignment for Mbp1 proteins and again color by conservation (easier to study) to briefly discuss the following question.
- Considering the conservation patterns for Mbp1 orthologues, and assuming that all these orthologues bind DNA in a similar way, which model appears to be more plausible for protein-DNA interactions in APSES domains? Is it the canonical, or the non-canonical binding mode? Discuss briefly what you would expect to find and how this relates to your observations. Distinguish clearly between experimental evidence, computational inference and empirical hypothesis. You are of course welcome to paste detail views (stereo !) of particular sidechains, or surfaces etc. if this helps your arguments. Sometimes a picture is worth many words. But this is not a requirement, we are more interested in evidence-based reasoning than in the form of the presentation.
(2 marks)
(5) Summary of Resources
- Links and background reading
- Review (PDF, restricted) Manuel Peitsch on Homology Modeling
- Review (PDF, restricted) Aravind et al. Helix-turn-helix domains
- Review (PDF, restricted) Gajiwala & Burley, winged-Helix domains
- Assigned Organisms
- PDB file format (see the Coordinate Section if you are unsure about chain identifiers)
- Wikipedia on Structural Superposition (although the article is called "Structural Alignment")
- Alignments
[End of assignment]
If you have any questions at all, don't hesitate to mail me at boris.steipe@utoronto.ca or post your question to the 2011 Course Mailing List .