Difference between revisions of "User:Boris/BCB hackathon 2018"
m (→Background) |
m (→Background) |
||
| Line 66: | Line 66: | ||
{{Smallvspace}} | {{Smallvspace}} | ||
| − | Our imagination of the genome has matured tremendously. '''What is the image we will find for | + | Our imagination of the genome has matured tremendously. |
| + | |||
| + | {{Smallvspace}} | ||
| + | |||
| + | '''What is the image we will find for the Human Genome – 20 years on?''' | ||
Revision as of 01:53, 7 February 2018
BCB Hackathon 2018
(Topic Proposal: The Human Genome - 20 years later)
Abstract:
This topic proposal for the 2018 BCB hackathon is to explore new ways to provide a holistic overview of the contents of the human genome, for the occasion of the 20th anniversary of its sequence.
Contents
Background
|
The first two draft sequences of the human genome were published in February of 2001[1][2]. Three years from now will mark the twentieth anniversary of this accomplishment that like now other has shaped the landscape of bioinformatics, computational biology and molecular medicine.
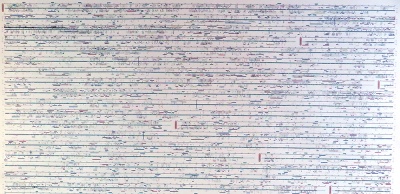
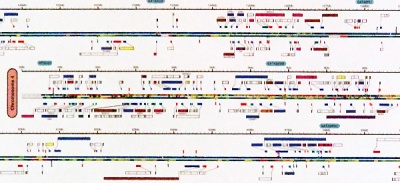
In 2001, Celera - a private company founded three years earlier to commercialize genome information - published an iconic poster summarizing their version of the genome. It is still fascinating today. |
|
|
This poster is significant, not so much for its interpretable content, but for the unique perspective it gives us on the entirety of information that constitutes our molecular identity. |
|
|
The details are rich, in fact, surprisingly "modern", presenting features like CpG islands and SNP density, and exon transcripts with Gene Ontology functional categories colour coded, for forward and reverse strand, accurately plotted on the nucleotide backbone at about 500 kB per centimetre. This was computed from gff records with Josep Abril's |
But we know so much more today. The number of sequenced genomes has exploded - selected individuals at first, then we envisioned the 1,000 genomes project (2008, completed 2012); quickly set our sights on 100,000 genomes (2012, almost completed), and as of today more than 500,000 human genomes have been sequenced overall. We have sequenced cancers, and genetic diseases. We have sequenced representatives of virtually all ethnicities on the planet. We have even sequenced Neanderthals and Denisovians, and we have sequenced other species far and wide to acquire a sense of where we fit into the landscape of evolution. We have annotated the contents of the genome in the ENCODE project. We have built databases that carefully dissect all proteins into their domains, such as InterPro. We have started to outline how things work together in functional networks such as the STRING data, or in modules as published by KEGG, and we are beginning to translate our insights into actionable information for medicine, at the OICR, at Sick Kids' TCAG.
Our imagination of the genome has matured tremendously.
What is the image we will find for the Human Genome – 20 years on?
Goals
Process
Perspectives
Notes
- ↑
Lander et al. (2001) Initial sequencing and analysis of the human genome. Nature 409:860-921. (pmid: 11237011) - ↑
Venter et al. (2001) The sequence of the human genome. Science 291:1304-51. (pmid: 11181995) - ↑
Abril & Guigó (2000) gff2ps: visualizing genomic annotations. Bioinformatics 16:743-4. (pmid: 11099262)
About ...
Last update:
- 2017-02-06
Version:
- 1.0
Version history:
- 1.0 First proposal
![]() This copyrighted material is licensed under a Creative Commons Attribution 4.0 International License. Follow the link to learn more.
This copyrighted material is licensed under a Creative Commons Attribution 4.0 International License. Follow the link to learn more.